| Motif ID |
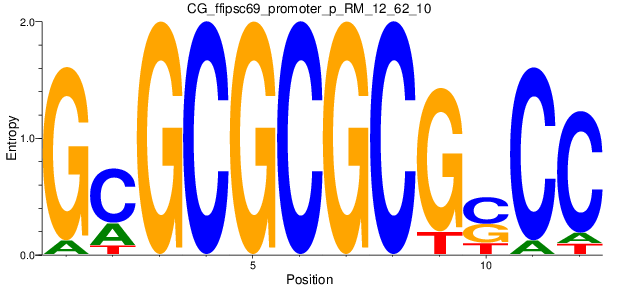
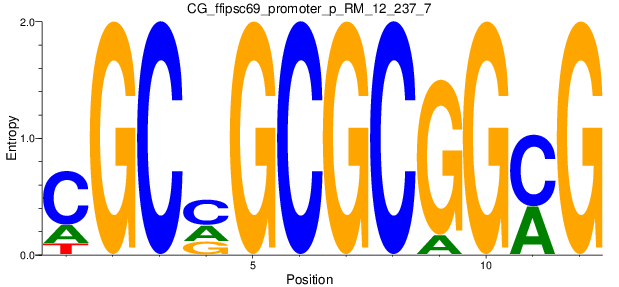
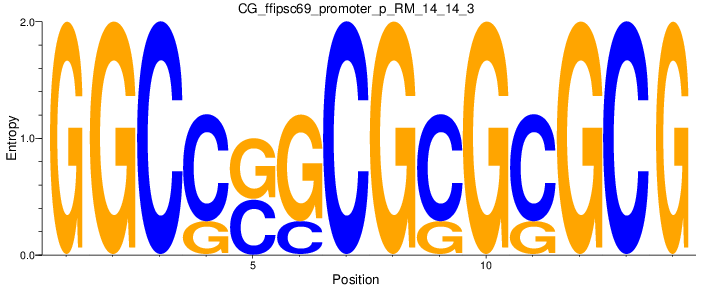
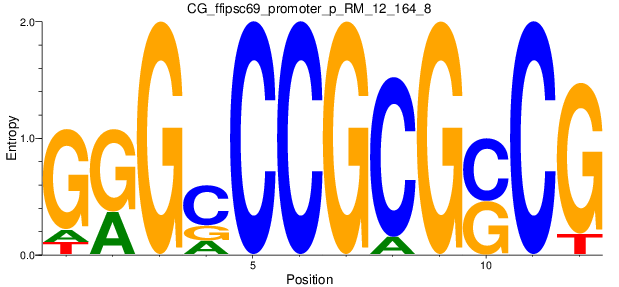
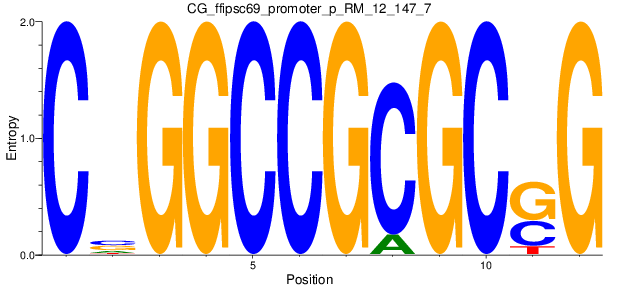
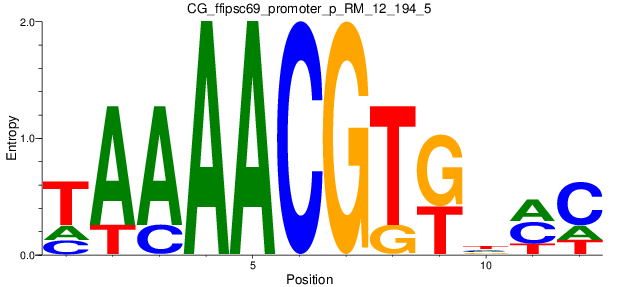
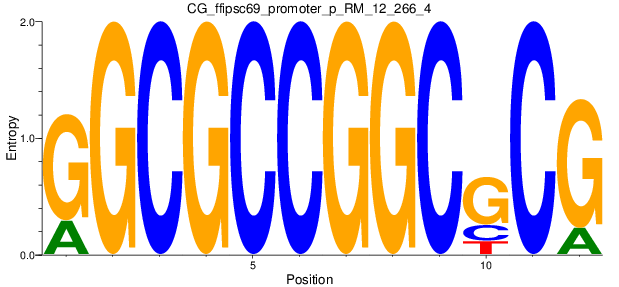
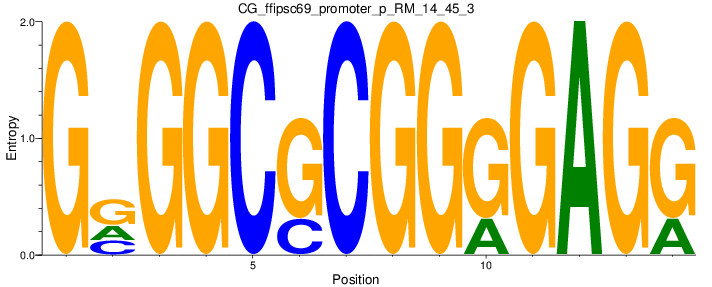
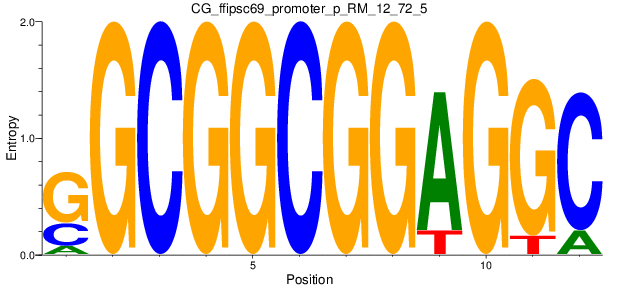
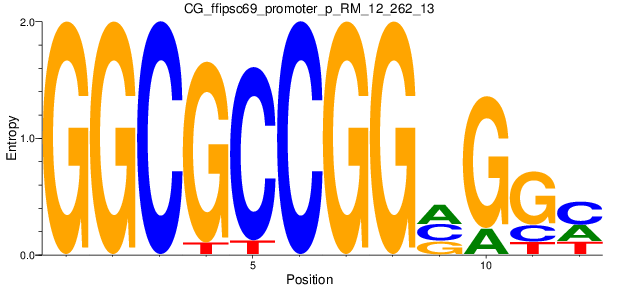
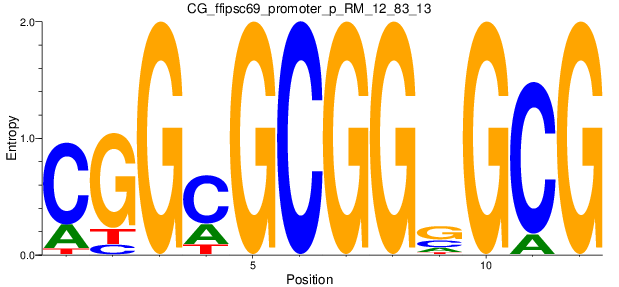
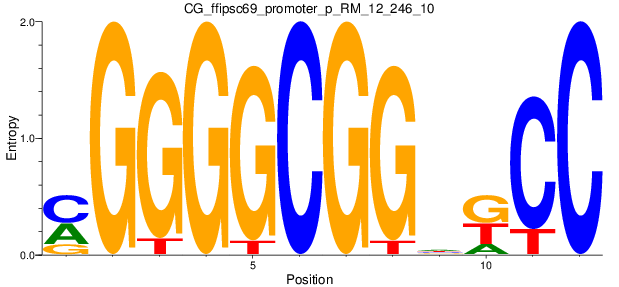
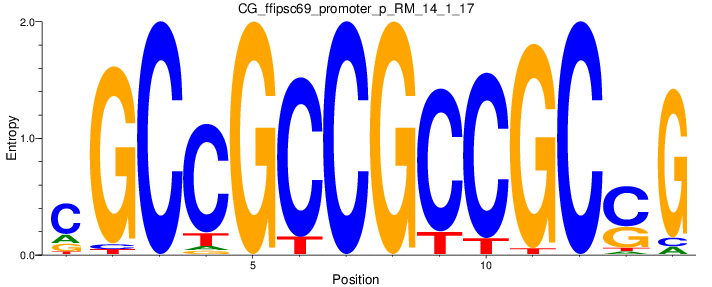
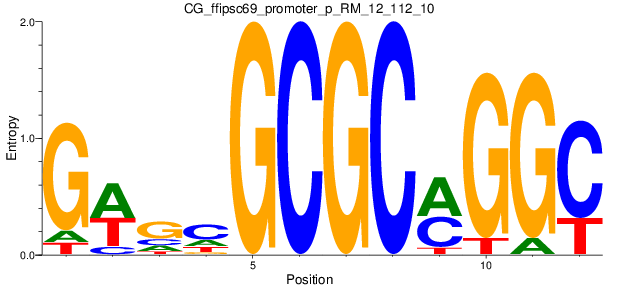
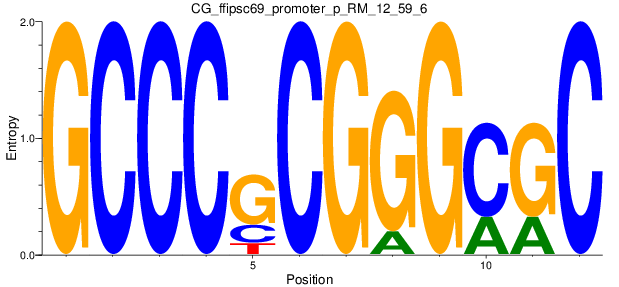
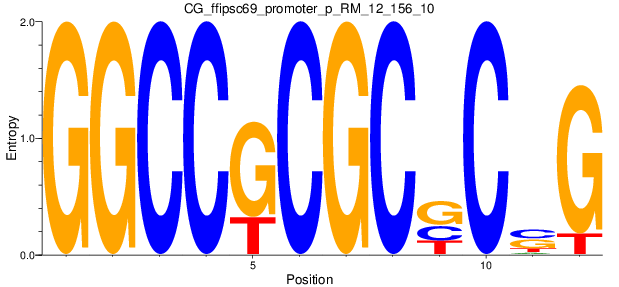
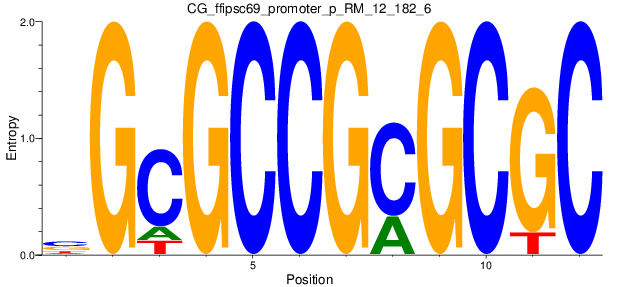
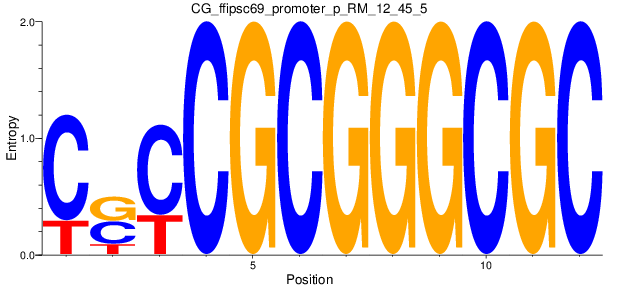
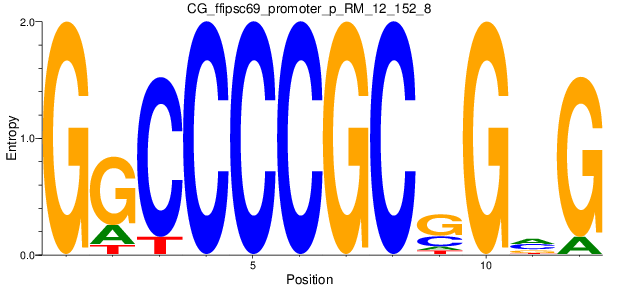
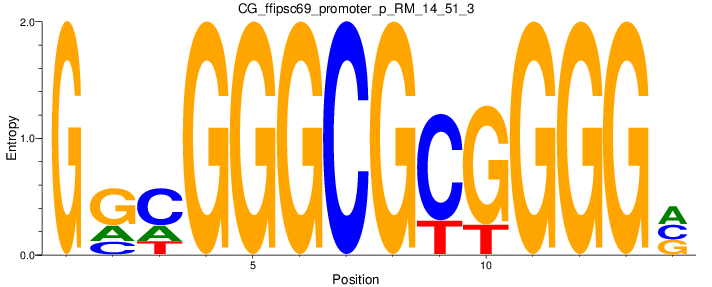
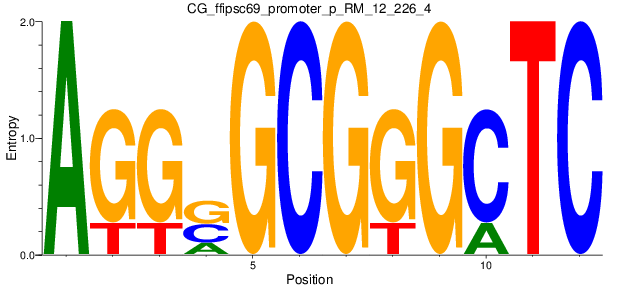
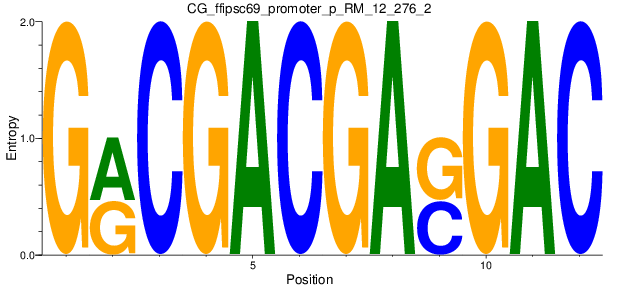
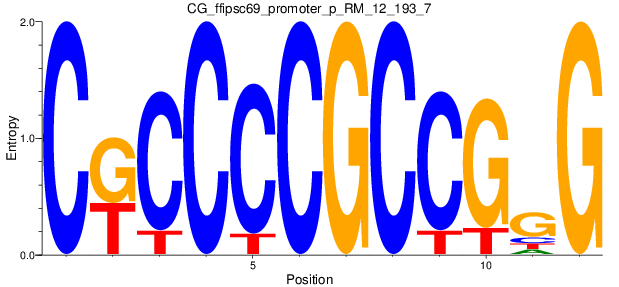
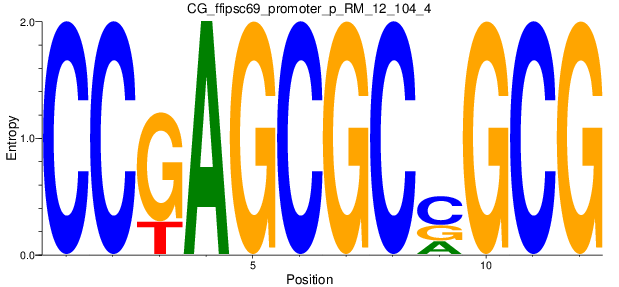
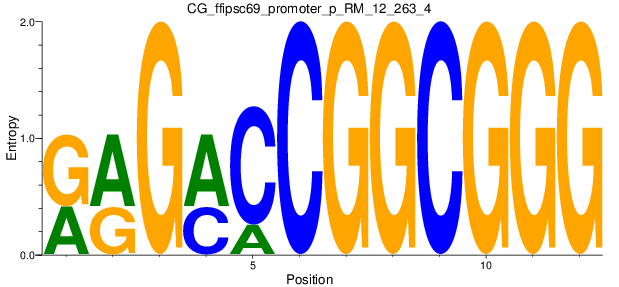
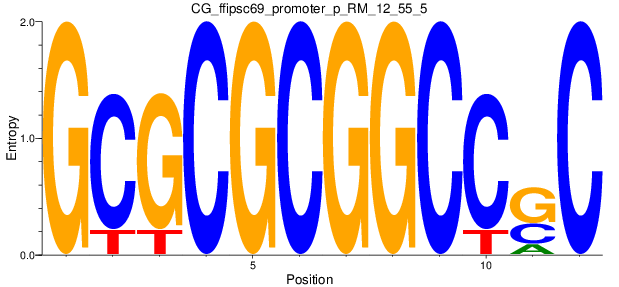
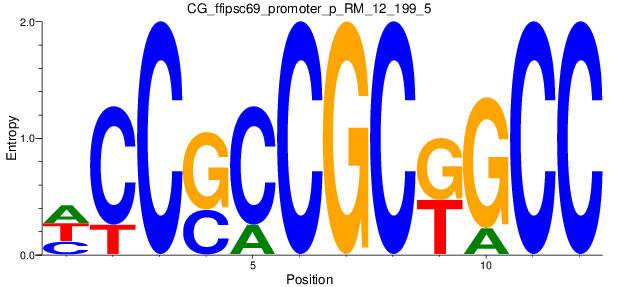
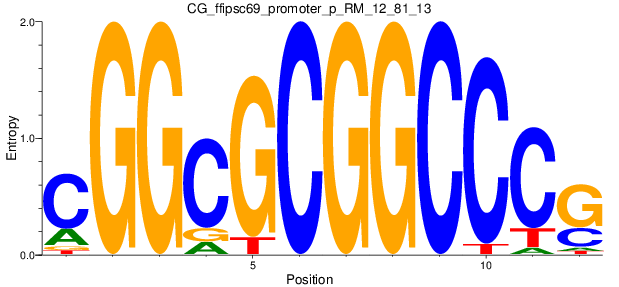
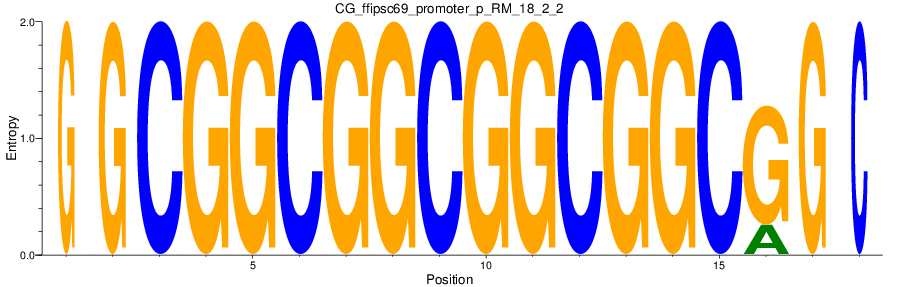
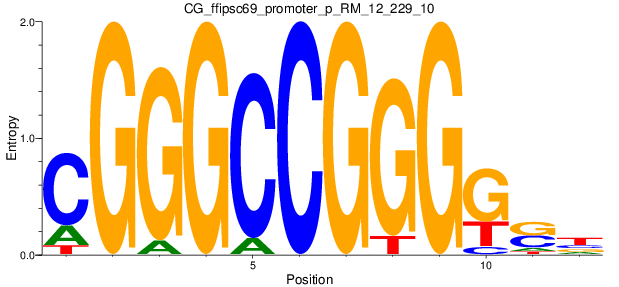
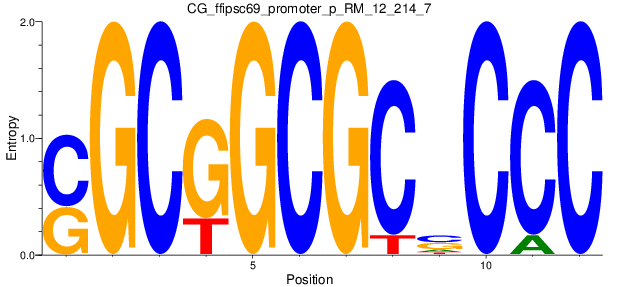
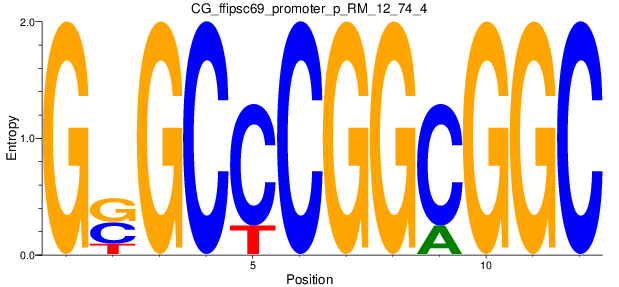
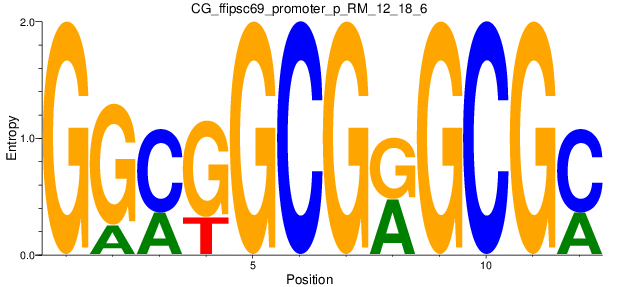
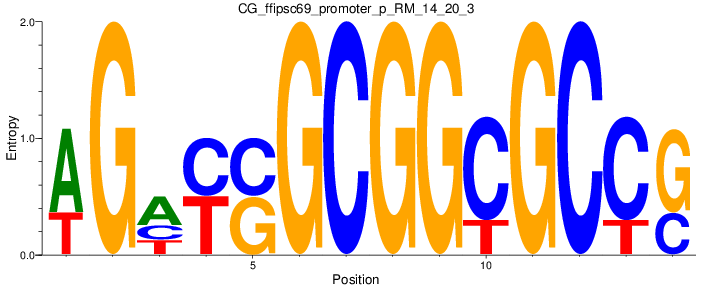
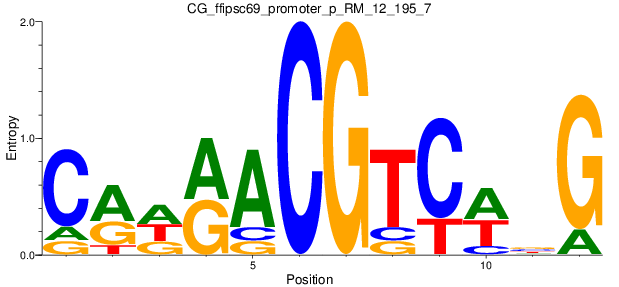
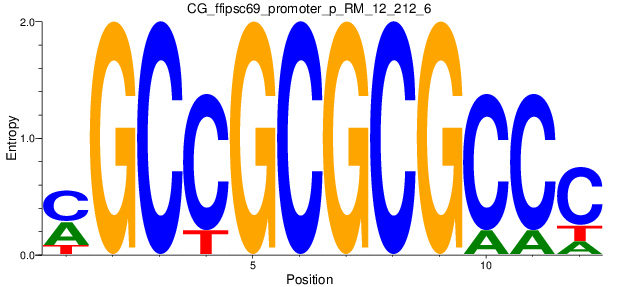
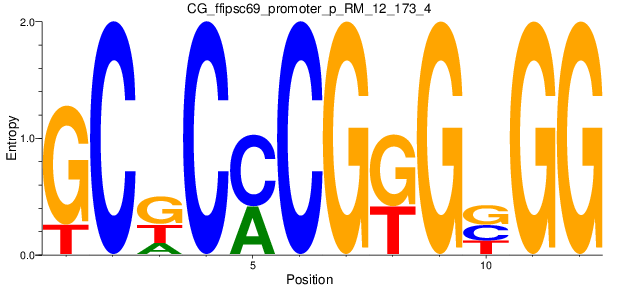
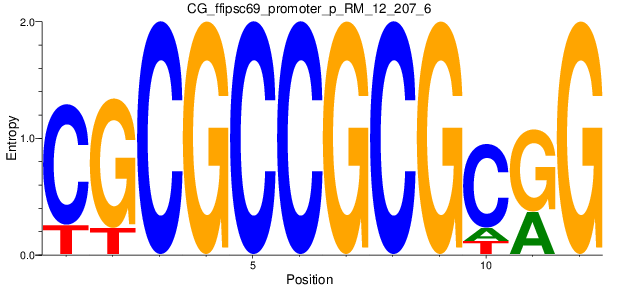
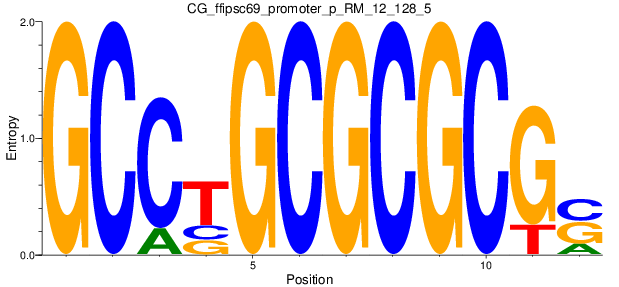
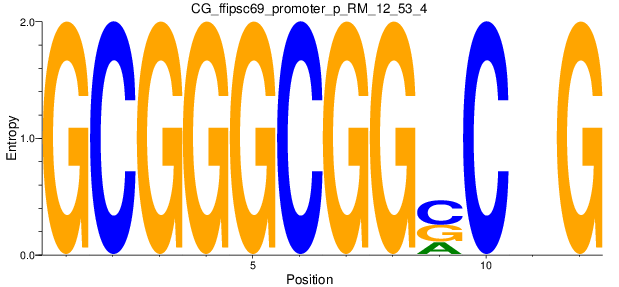
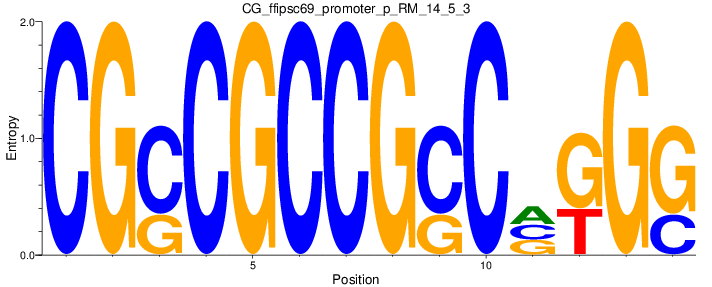
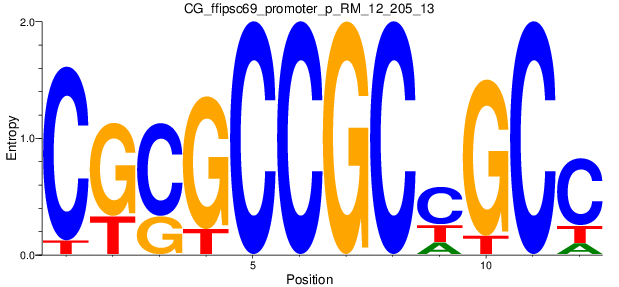
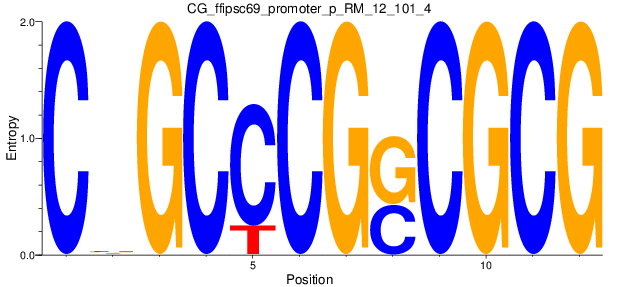
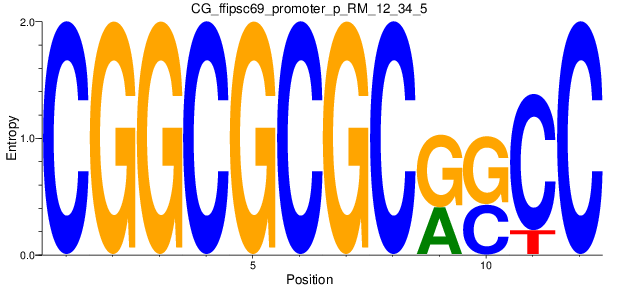
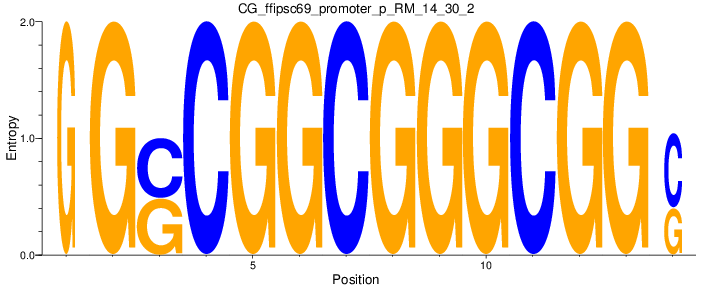
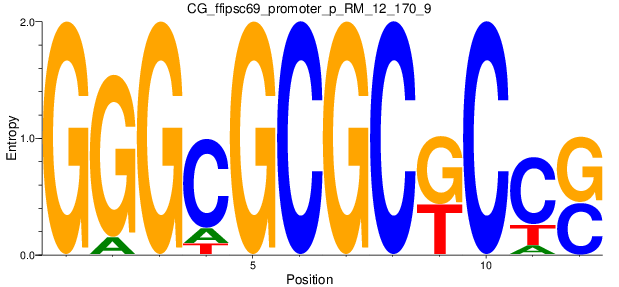
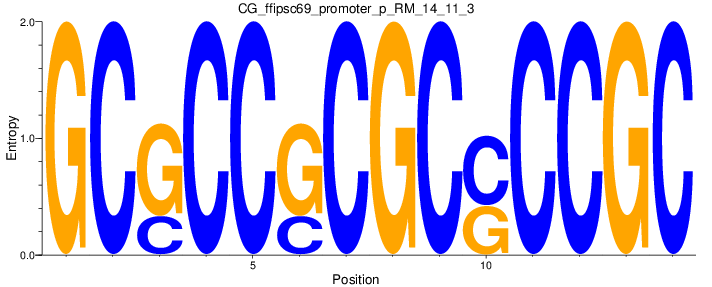
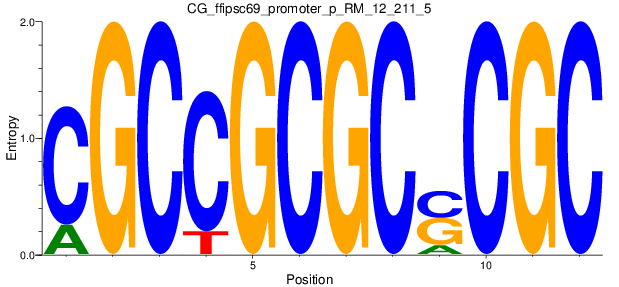
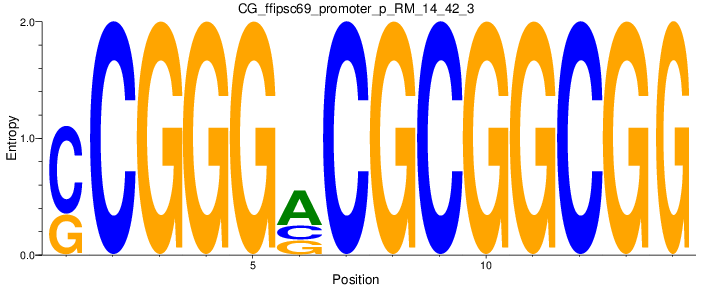
Motif Logo |
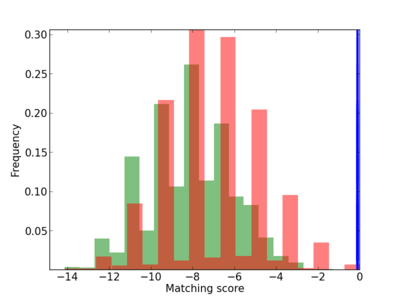
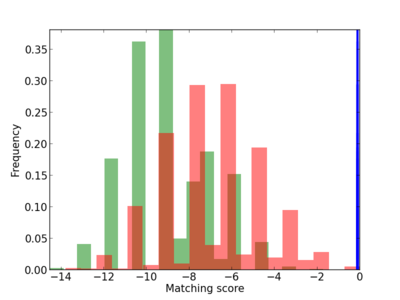
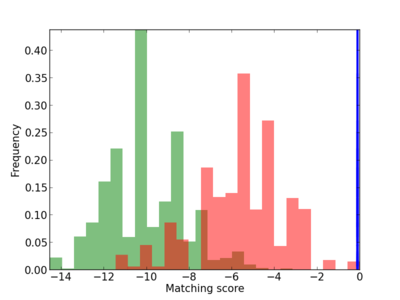
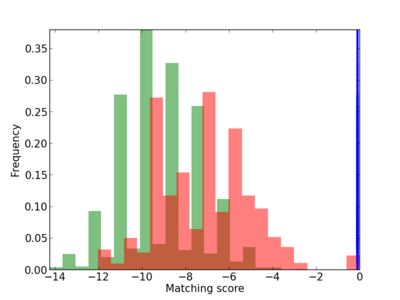
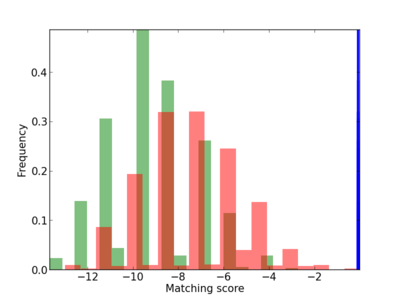
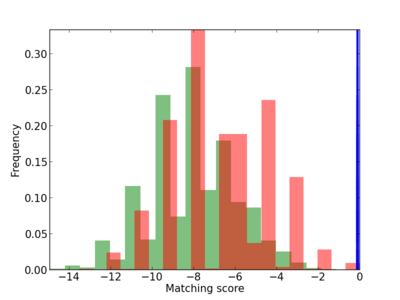
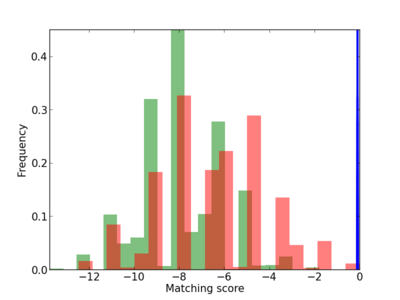
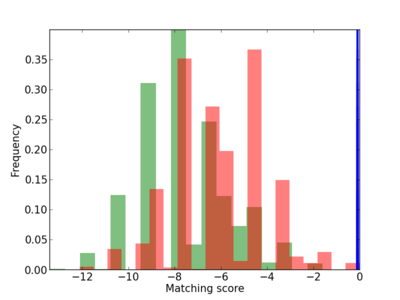
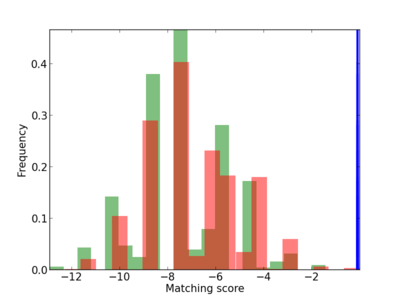
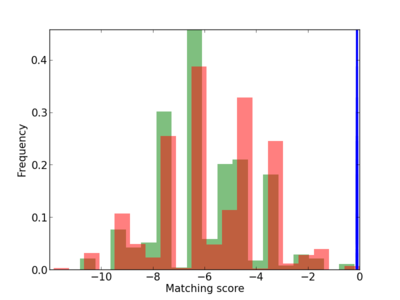
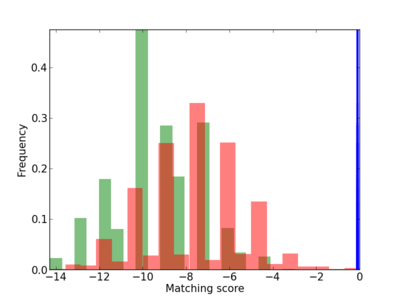
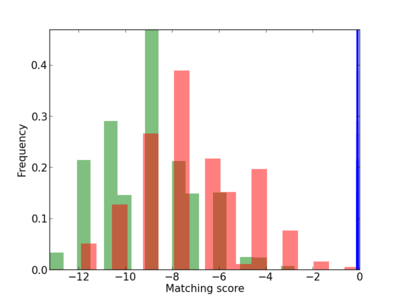
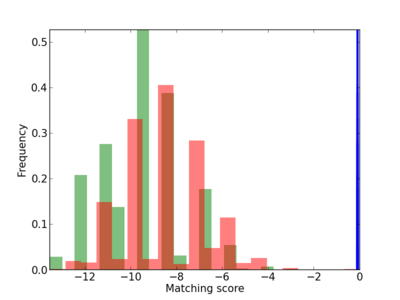
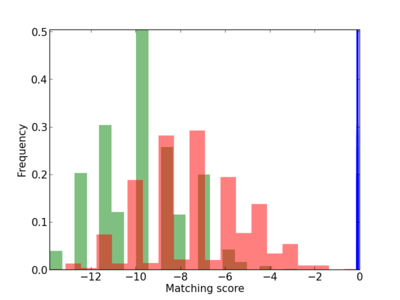
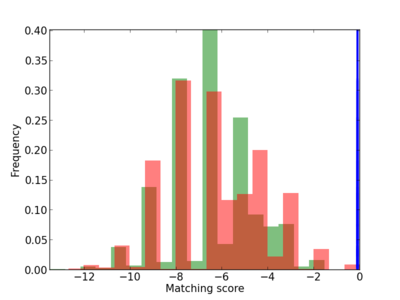
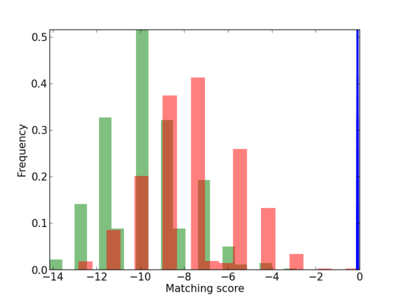
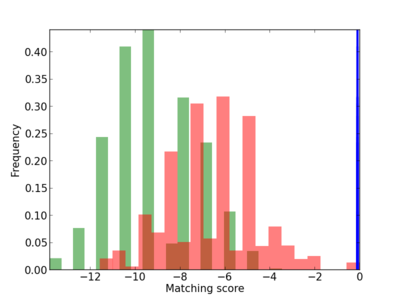
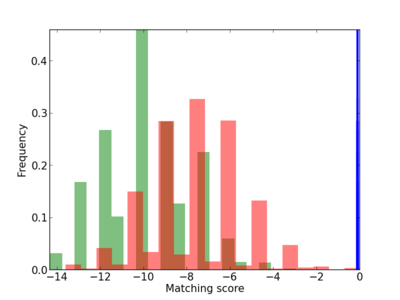
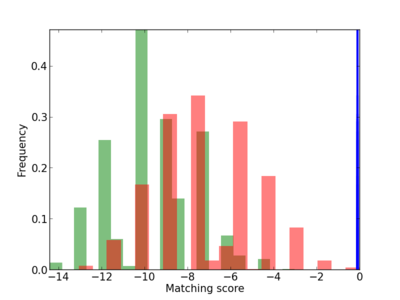
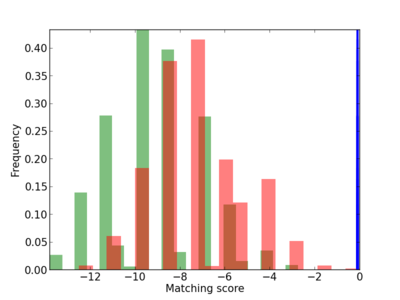
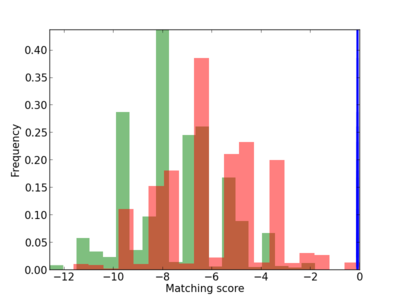
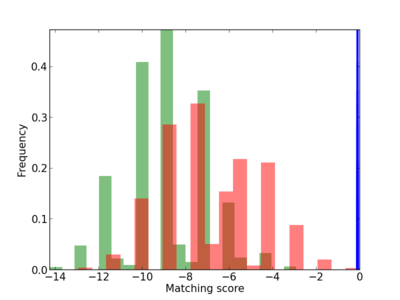
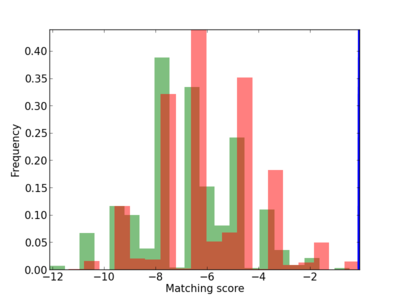
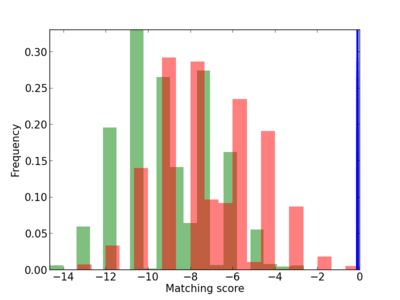
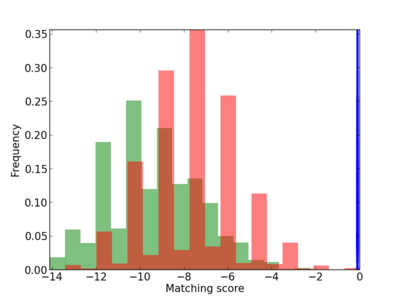
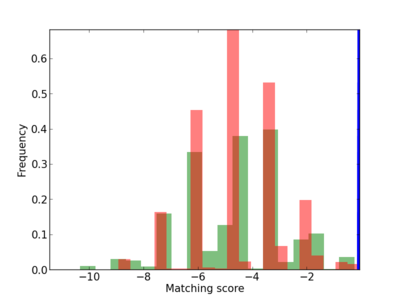
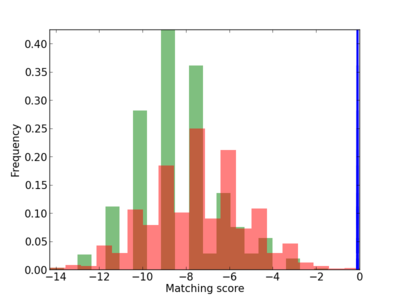
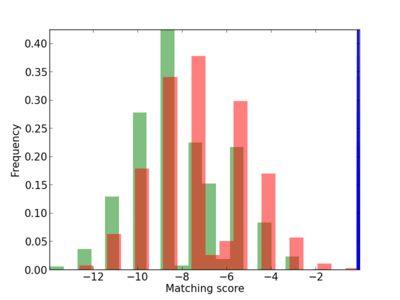
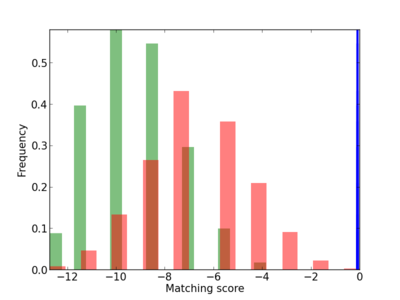
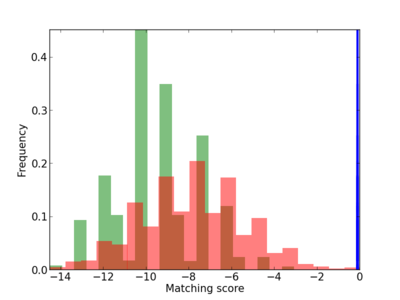
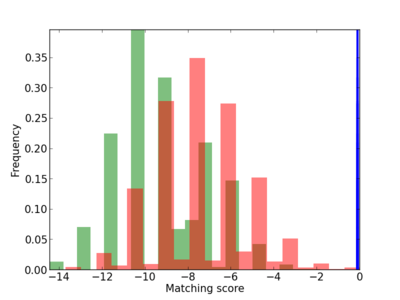
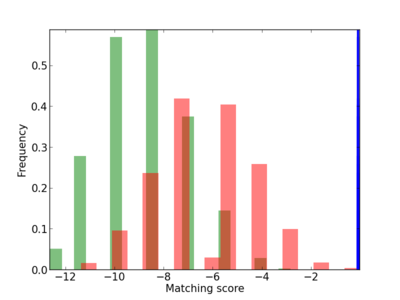
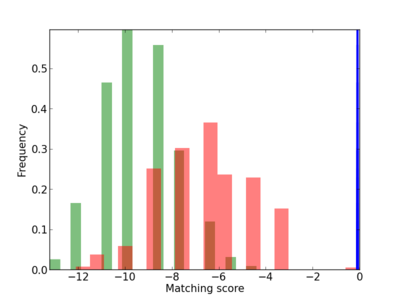
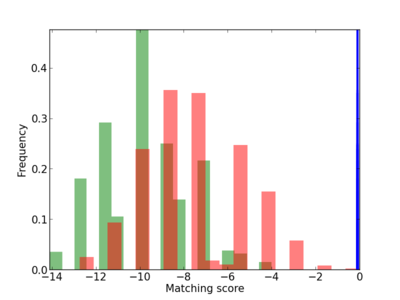
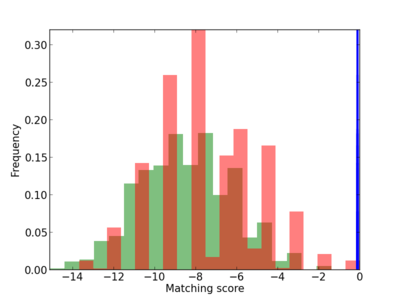
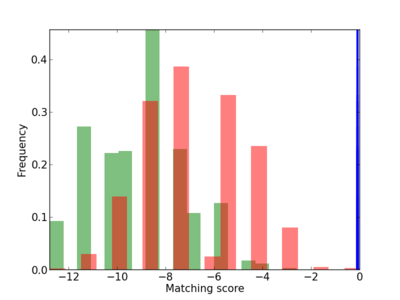
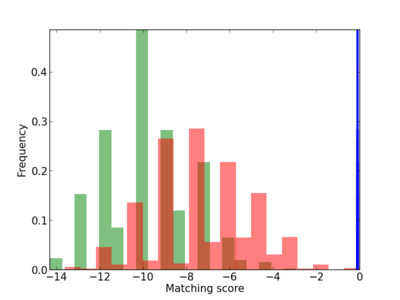
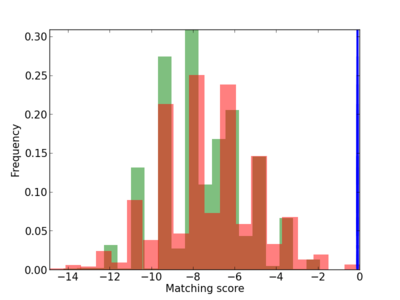
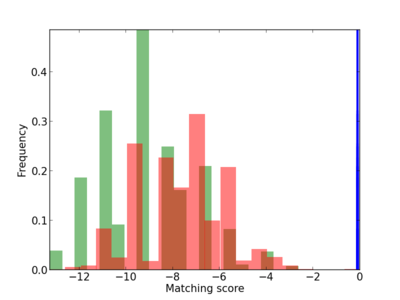
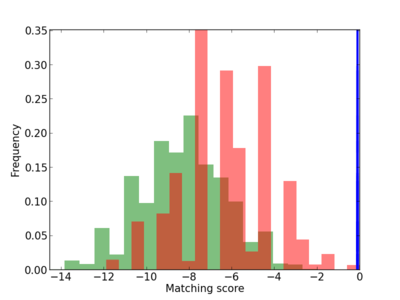
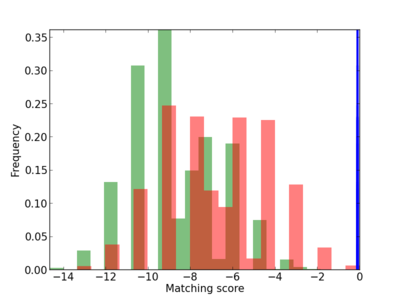
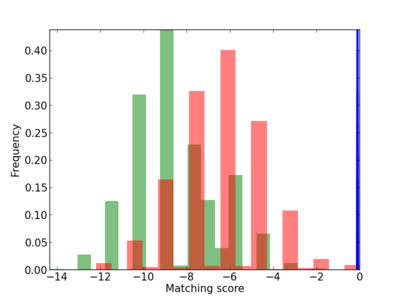
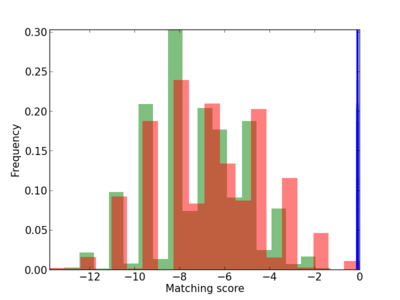
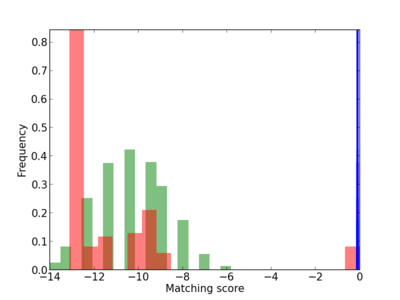
Scanned Score Distribution |
Frequency of Motif |
Replicability Score |
Evolutionary Conservation Score |
Gene Targets |
Gene Ontology of CpGMM targets |
| CG_ffipsc69_promoter_p_RM_12_62_10 |
 |
 |
5 |
0.8 |
3.65E-04 |
MAVS |
positive regulation of IP-10 production [BP] |
| CG_ffipsc69_promoter_p_RM_12_237_7 |
 |
 |
4 |
0.6 |
3.29E-04 |
TGIF1 |
nodal signaling pathway [BP] |
| CG_ffipsc69_promoter_p_RM_12_58_12 |
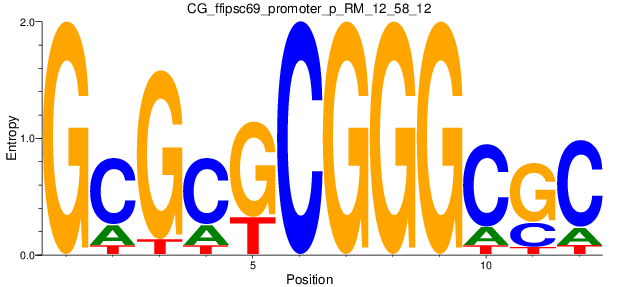 |
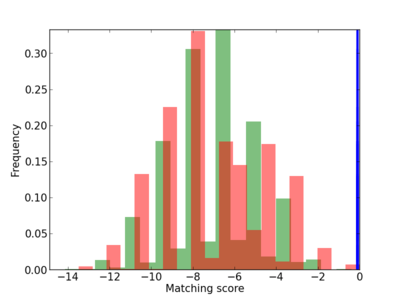 |
5 |
0.8 |
1.19E-04 |
OPRL1,
BIRC7 |
nociceptin receptor activity [MF],
regulation of natural killer cell apoptotic process [BP] |
| CG_ffipsc69_promoter_p_RM_12_159_6 |
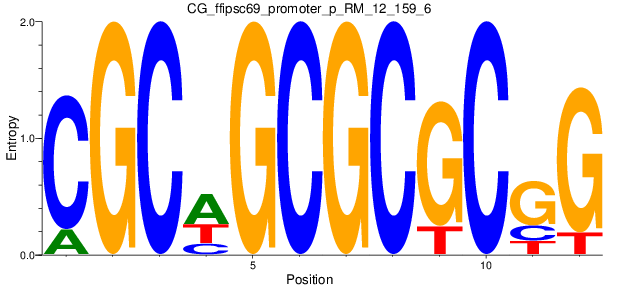 |
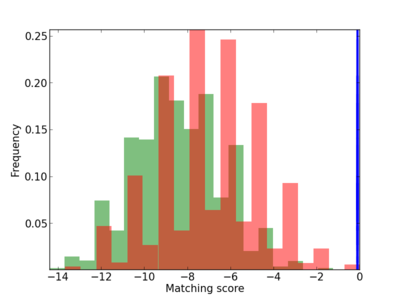 |
5 |
0.8 |
2.75E-04 |
TPM1,
SRD5A3 |
positive regulation of heart rate by epinephrine [BP],
polyprenol catabolic process [BP] |
| CG_ffipsc69_promoter_p_RM_12_204_6 |
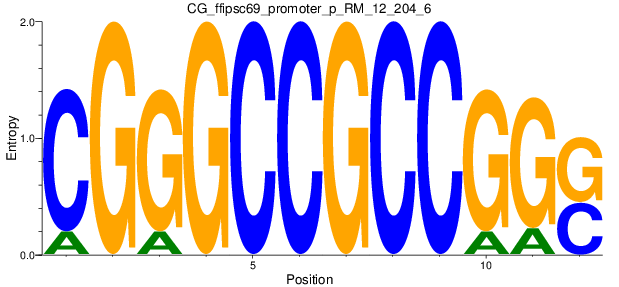 |
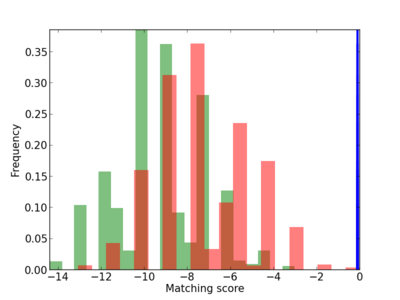 |
5 |
1.0 |
3.10E-04 |
FPGS,
SMAD4 |
transforming growth factor beta receptor, common-partner cytoplasmic mediator activity [MF],
tetrahydrofolylpolyglutamate synthase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_290_7 |
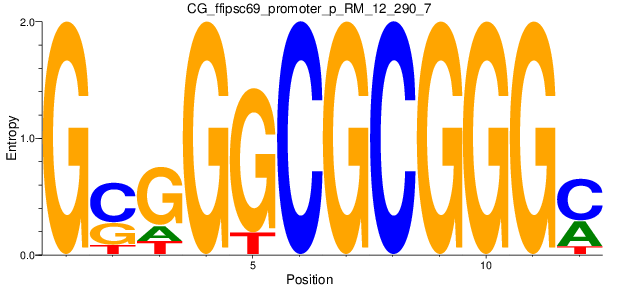 |
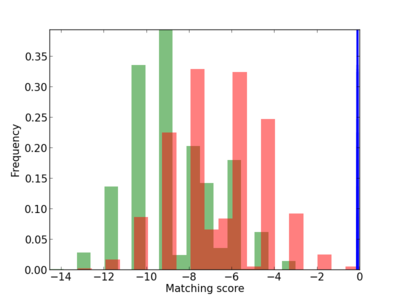 |
5 |
1.0 |
1.28E-04 |
PNRC2 |
deadenylation-independent decapping of nuclear-transcribed mRNA [BP] |
| CG_ffipsc69_promoter_p_RM_14_16_1 |
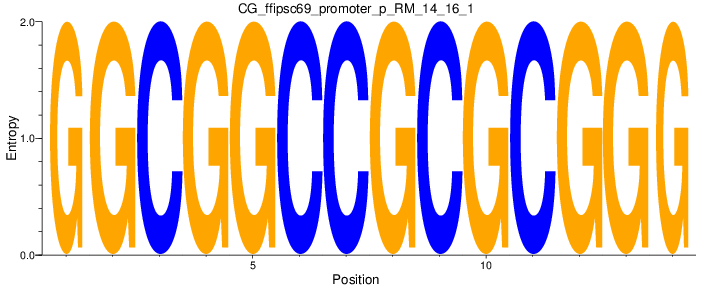 |
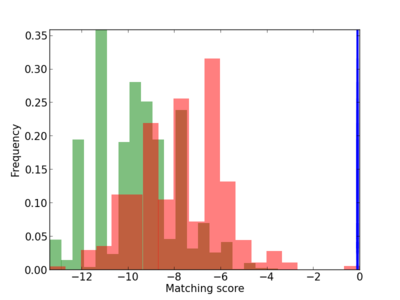 |
3 |
0.4 |
1.33E-04 |
PKD2 |
metanephric smooth muscle tissue development [BP],
HLH domain binding [MF],
detection of nodal flow [BP],
metanephric cortex development [BP],
metanephric ascending thin limb development [BP],
polycystin complex [CC],
cellular response to hydrostatic pressure [BP],
positive regulation of inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity [BP],
calcium-induced calcium release activity [MF],
renal artery morphogenesis [BP],
metanephric cortical collecting duct development [BP],
basal cortex [CC] |
| CG_ffipsc69_promoter_p_RM_12_13_10 |
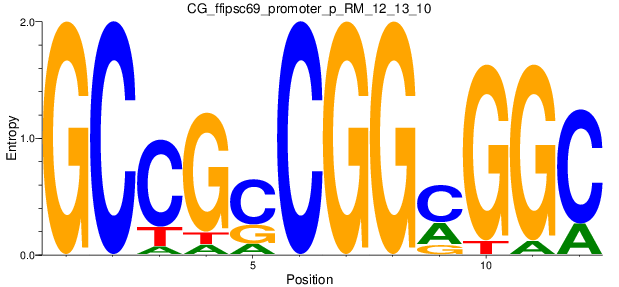 |
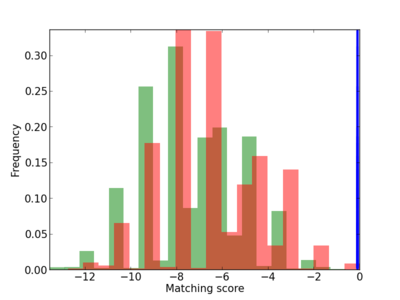 |
4 |
0.6 |
3.99E-04 |
RAB10 |
establishment of protein localization to endoplasmic reticulum membrane [BP] |
| CG_ffipsc69_promoter_p_RM_12_157_3 |
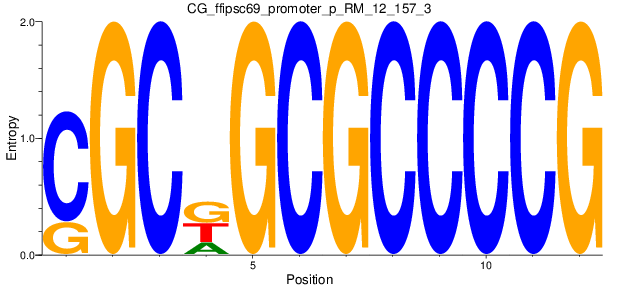 |
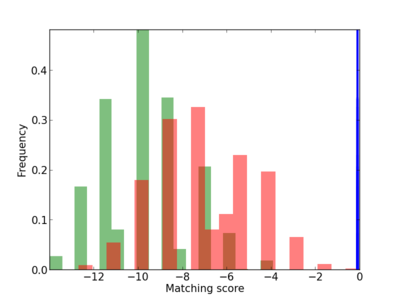 |
2 |
0.4 |
2.36E-04 |
PRDX5 |
peroxynitrite reductase activity [MF] |
| CG_ffipsc69_promoter_p_RM_14_43_3 |
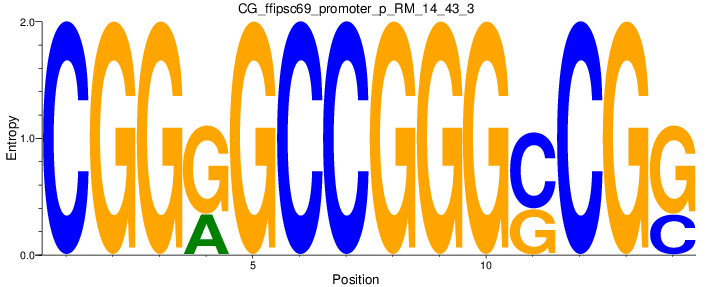 |
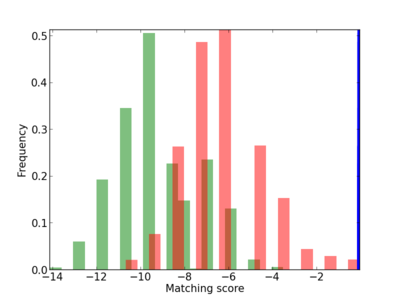 |
4 |
0.6 |
2.75E-04 |
MACF1,
ALDH1A3,
PPARD |
linoleic acid binding [MF],
posttranslational protein targeting to membrane [BP],
nucleus accumbens development [BP] |
| CG_ffipsc69_promoter_p_RM_12_225_5 |
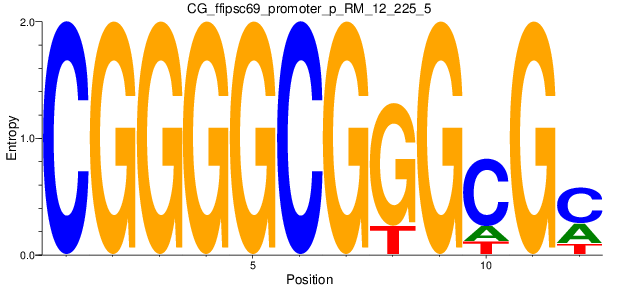 |
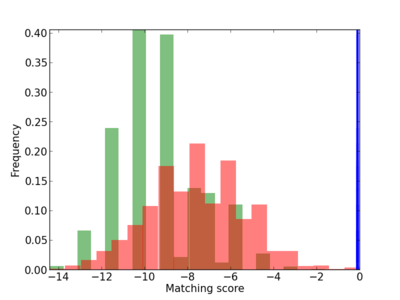 |
3 |
0.6 |
1.97E-04 |
TALDO1,
GLYCTK |
sedoheptulose-7-phosphate:D-glyceraldehyde-3-phosphate glyceronetransferase activity [MF],
glycerate kinase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_122_12 |
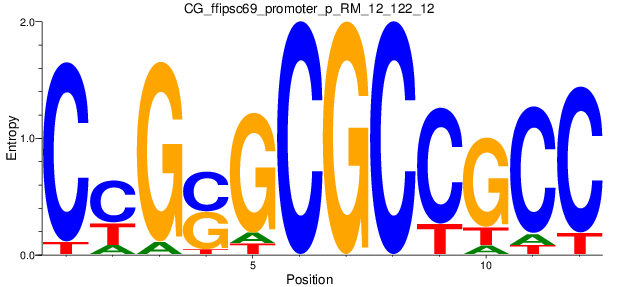 |
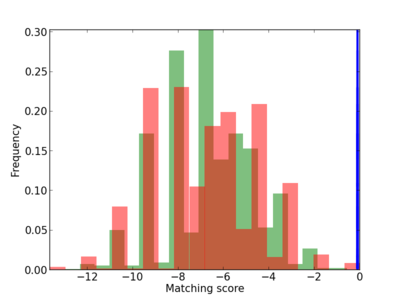 |
3 |
0.6 |
2.63E-04 |
AP2M1,
EXOSC2,
HMGA2,
AP2A2,
ISG20 |
clathrin coat of coated pit [CC],
response to virus [BP],
3'-5'-exoribonuclease activity [MF],
regulation of defense response to virus by virus [BP],
clathrin-coated endocytic vesicle membrane [CC],
clathrin adaptor complex [CC] |
| CG_ffipsc69_promoter_p_RM_12_98_7 |
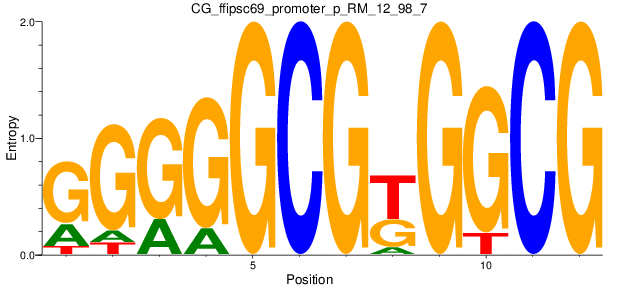 |
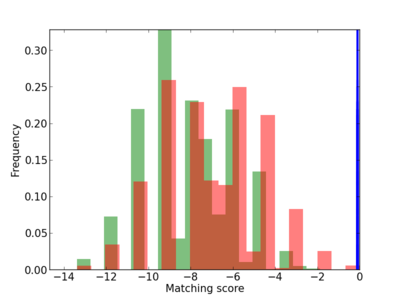 |
4 |
0.6 |
3.25E-04 |
MCCC2 |
coenzyme A metabolic process [BP] |
| CG_ffipsc69_promoter_p_RM_12_223_9 |
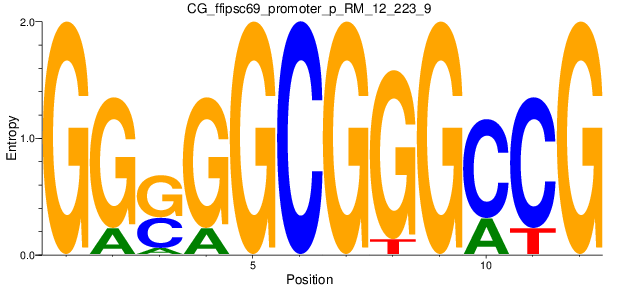 |
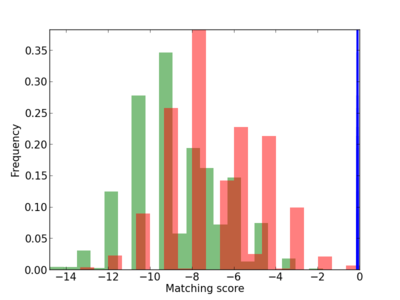 |
4 |
0.6 |
2.33E-04 |
PRDX5,
ITGAV,
LMNA |
regulation of apoptosis involved in tissue homeostasis [BP],
negative regulation of entry of bacterium into host cell [BP],
sterol regulatory element binding protein import into nucleus [BP],
entry of symbiont into host cell by promotion of host phagocytosis [BP] |
| CG_ffipsc69_promoter_p_RM_14_14_3 |
 |
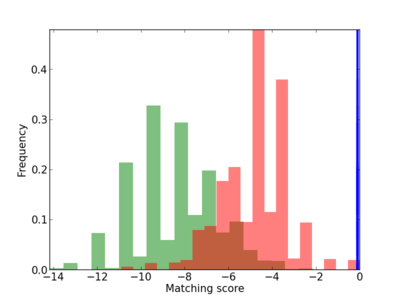 |
2 |
0.4 |
4.78E-04 |
UBC,
DOLPP1 |
hypothalamus gonadotrophin-releasing hormone neuron development [BP],
dolichyldiphosphatase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_256_3 |
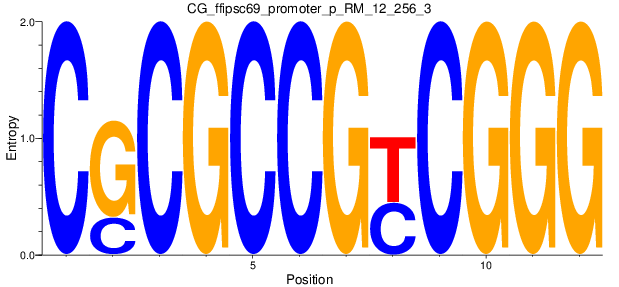 |
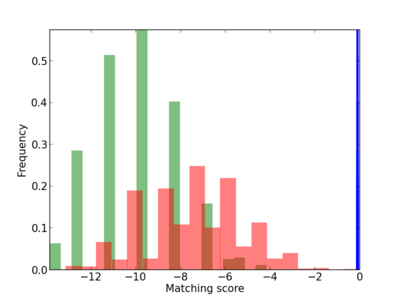 |
4 |
0.8 |
4.04E-04 |
HEY2,
STAT5A |
positive regulation of mast cell differentiation [BP],
negative regulation of cardiac vascular smooth muscle cell differentiation [BP] |
| CG_ffipsc69_promoter_p_RM_12_252_5 |
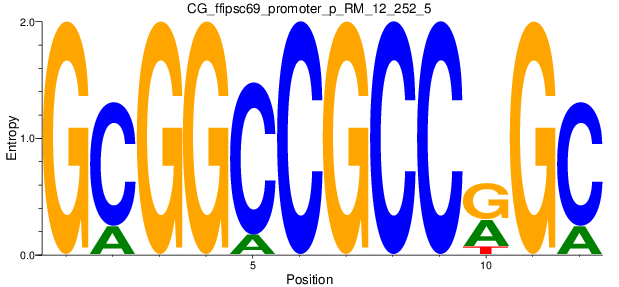 |
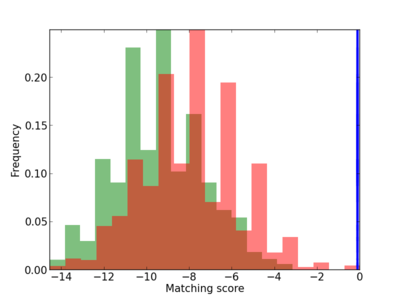 |
2 |
0.4 |
3.79E-04 |
PIWIL4,
NCBP1,
ECT2,
PRPF18,
COMMD1,
HOXC4,
NUP188,
ZNF583,
AEBP1,
ALKBH5,
CSRP2BP,
SYCP2L |
mRNA transport [BP],
nucleus [CC] |
| CG_ffipsc69_promoter_p_RM_12_264_5 |
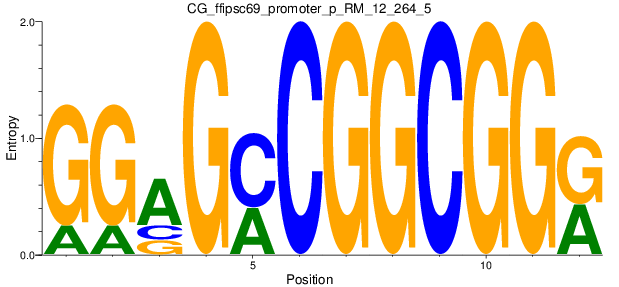 |
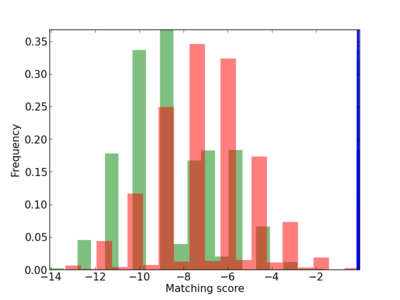 |
5 |
0.6 |
3.95E-04 |
UAP1,
WEE2 |
UDP-N-acetylglucosamine diphosphorylase activity [MF],
negative regulation of oocyte development [BP],
female pronucleus assembly [BP],
meiotic metaphase II [BP] |
| CG_ffipsc69_promoter_p_RM_12_166_10 |
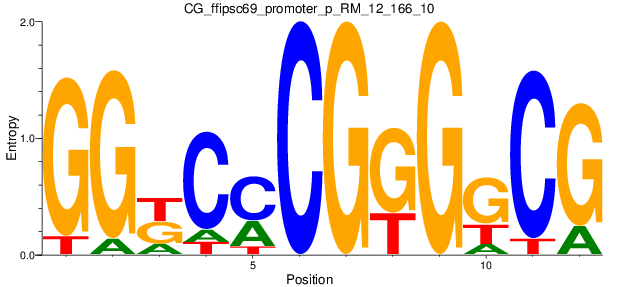 |
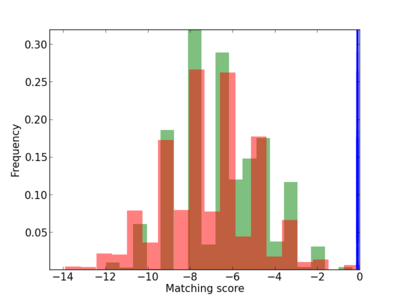 |
4 |
0.8 |
3.25E-05 |
F2RL1,
DOLPP1 |
negative regulation of toll-like receptor 3 signaling pathway [BP],
positive regulation of cytokine secretion involved in immune response [BP],
dolichyl diphosphate biosynthetic process [BP],
positive regulation of eosinophil degranulation [BP],
dolichyldiphosphatase activity [MF],
positive regulation of toll-like receptor 2 signaling pathway [BP],
chemokine (C-C motif) ligand 2 secretion [BP],
positive regulation of neutrophil mediated killing of gram-negative bacterium [BP],
negative regulation of chemokine secretion [BP],
positive regulation of renin secretion into blood stream [BP] |
| CG_ffipsc69_promoter_p_RM_12_162_7 |
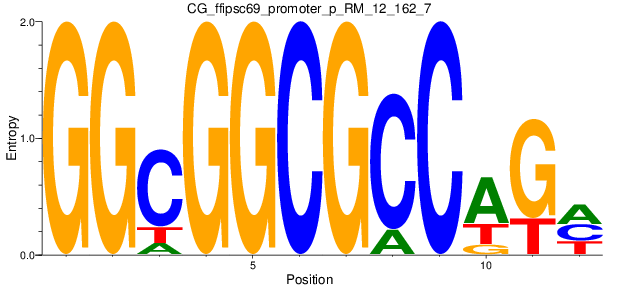 |
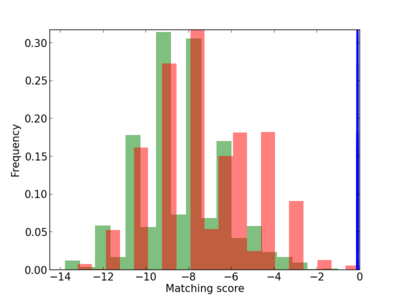 |
6 |
1.0 |
4.00E-04 |
EID1 |
histone acetyltransferase regulator activity [MF] |
| CG_ffipsc69_promoter_p_RM_14_29_3 |
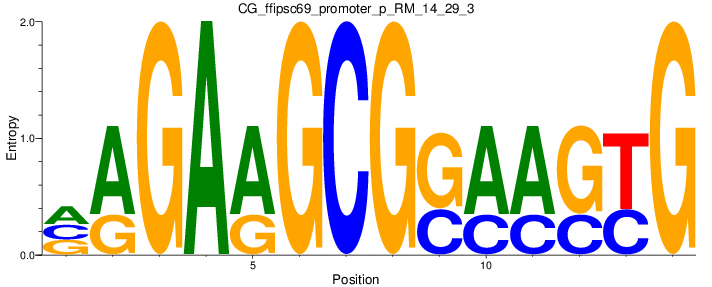 |
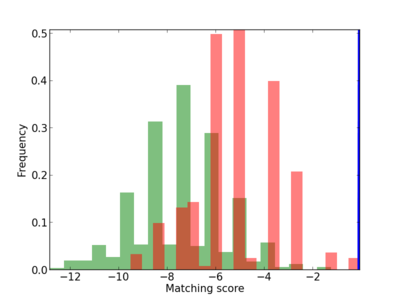 |
2 |
0.4 |
4.99E-04 |
HIST3H2BB,
RPS21,
HIST1H2BO |
nucleosome assembly [BP],
nucleosome [CC],
endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) [BP] |
| CG_ffipsc69_promoter_p_RM_12_164_8 |
 |
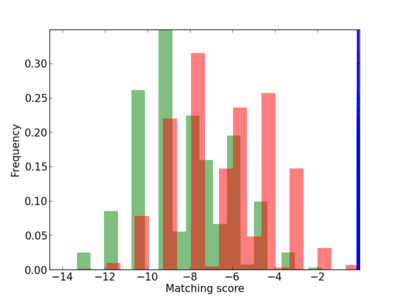 |
4 |
0.8 |
2.90E-04 |
MEIS1,
FOXE1 |
lens morphogenesis in camera-type eye [BP] |
| CG_ffipsc69_promoter_p_RM_12_147_7 |
 |
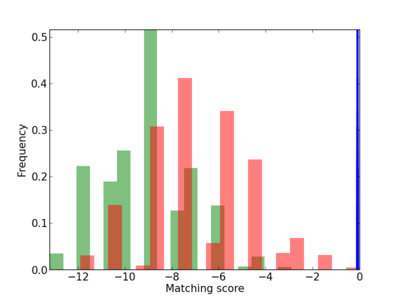 |
4 |
0.8 |
2.89E-04 |
HOXD1,
CUX2,
TELO2,
IRX1,
TGIF1,
RPTOR |
sequence-specific DNA binding [MF],
TORC1 complex [CC] |
| CG_ffipsc69_promoter_p_RM_12_126_5 |
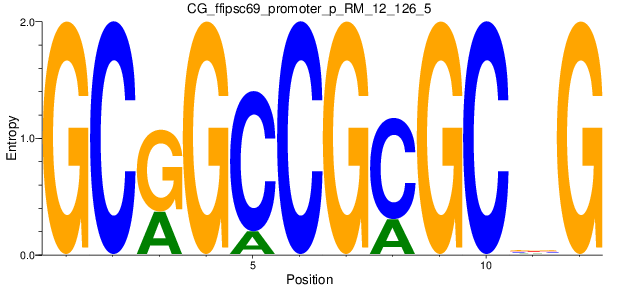 |
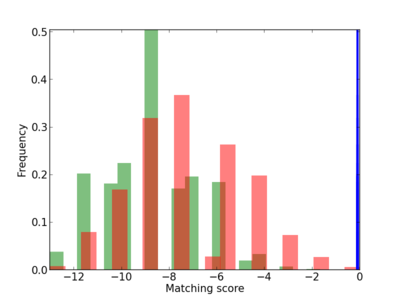 |
6 |
1.0 |
2.87E-04 |
BCL2L11,
PTF1A,
NOG,
LIMD1,
MTA1,
MAPK8IP3,
VWC2 |
negative regulation of BMP signaling pathway [BP],
embryo development [BP],
negative regulation of osteoblast differentiation [BP],
in utero embryonic development [BP],
signal transduction [BP] |
| CG_ffipsc69_promoter_p_RM_14_41_3 |
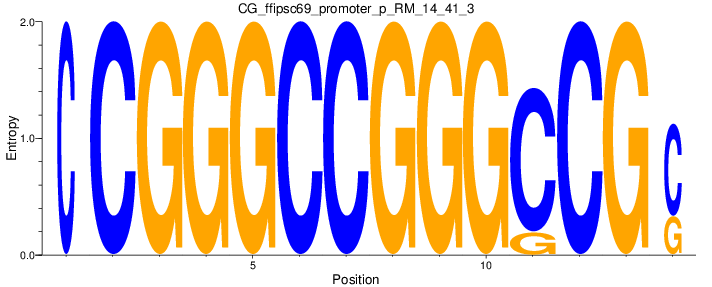 |
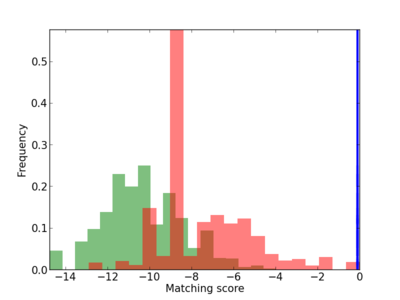 |
4 |
0.6 |
5.58E-04 |
MAN1B1 |
endoplasmic reticulum quality control compartment [CC] |
| CG_ffipsc69_promoter_p_RM_12_295_6 |
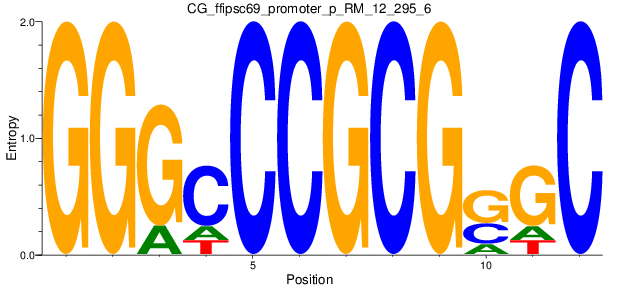 |
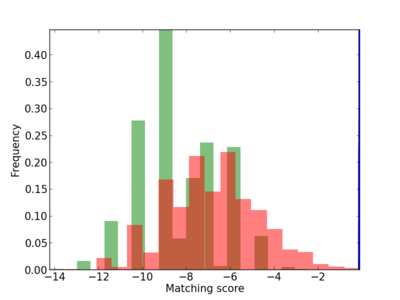 |
4 |
0.8 |
1.83E-04 |
EVX1 |
regulation of transcription from RNA polymerase II promoter involved in ventral spinal cord interneuron specification [BP] |
| CG_ffipsc69_promoter_p_RM_12_192_6 |
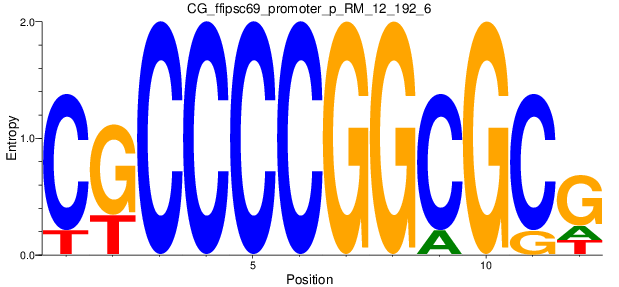 |
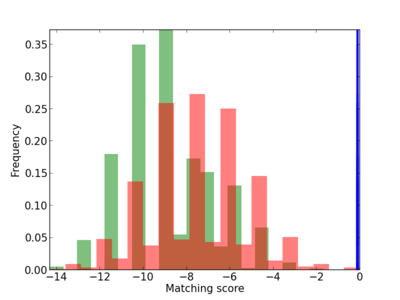 |
3 |
0.6 |
2.45E-04 |
PIK3CD,
STRA8,
PDPK1 |
3-phosphoinositide-dependent protein kinase activity [MF],
female meiosis sister chromatid cohesion [BP],
mast cell degranulation [BP],
T cell receptor signaling pathway [BP],
meiotic DNA double-strand break formation [BP],
meiotic cell cycle DNA replication checkpoint [BP] |
| CG_ffipsc69_promoter_p_RM_12_141_5 |
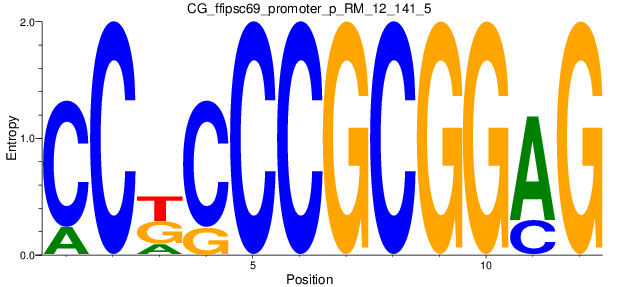 |
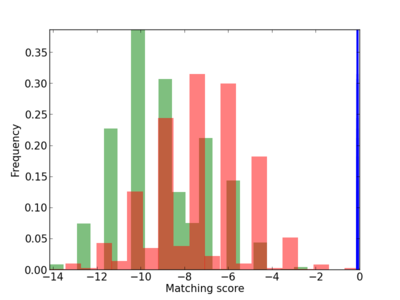 |
3 |
0.6 |
1.99E-04 |
CTCF |
chromatin insulator sequence binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_149_8 |
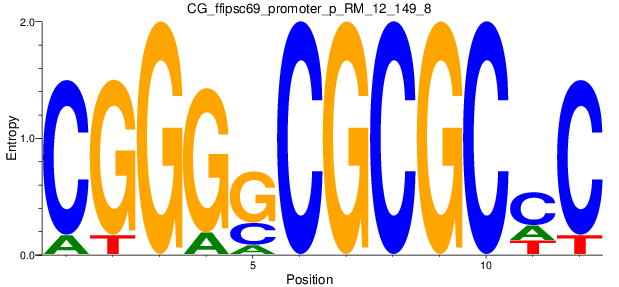 |
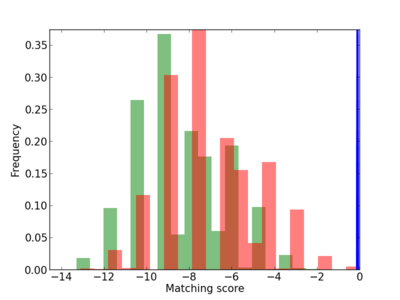 |
2 |
0.4 |
2.64E-04 |
ACADS,
TPI1 |
butyryl-CoA dehydrogenase activity [MF],
triose-phosphate isomerase activity [MF] |
| CG_ffipsc69_promoter_p_RM_14_31_4 |
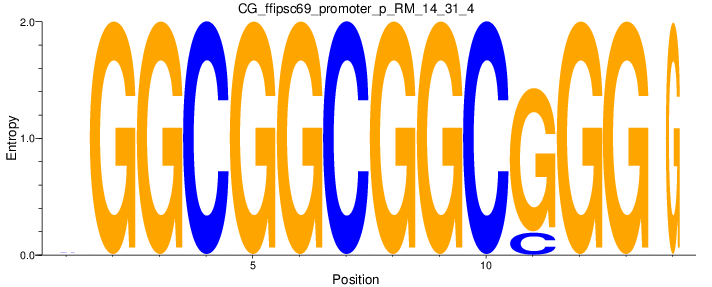 |
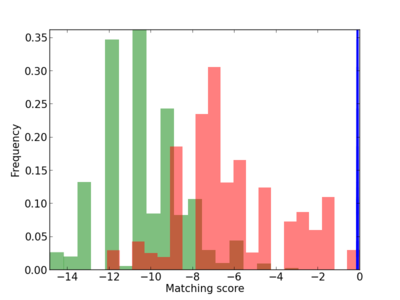 |
4 |
0.6 |
4.98E-04 |
RNF146,
MYO1B |
poly-ADP-D-ribose binding [MF],
actin filament-based movement [BP] |
| CG_ffipsc69_promoter_p_RM_12_52_3 |
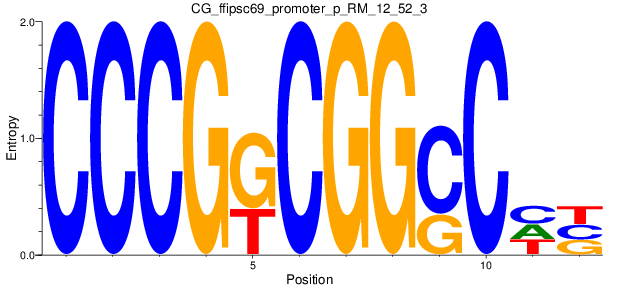 |
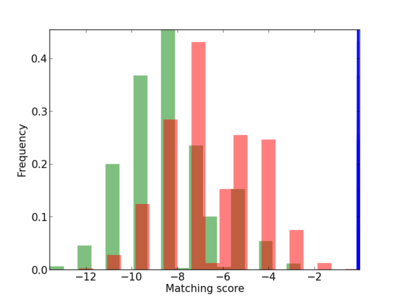 |
4 |
0.8 |
1.78E-04 |
ACAD10,
GOSR1,
CDC42,
IVD |
isovaleryl-CoA dehydrogenase activity [MF],
actin filament branching [BP],
intra-Golgi vesicle-mediated transport [BP],
flavin adenine dinucleotide binding [MF],
acyl-CoA dehydrogenase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_194_5 |
 |
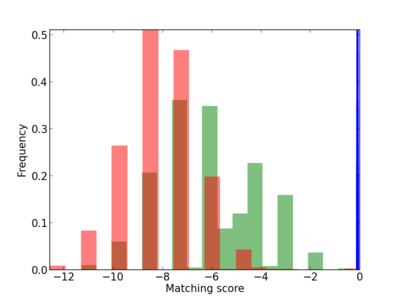 |
1 |
0.2 |
7.77E-05 |
TPST1,
ENGASE |
peptidyl-tyrosine sulfation [BP],
mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase activity [MF],
protein-tyrosine sulfotransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_266_4 |
 |
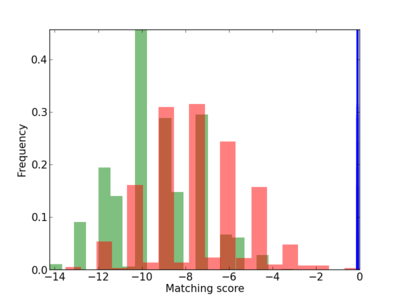 |
3 |
0.6 |
2.34E-04 |
GATA6,
SMAD6,
CEPT1,
MMP14 |
response to estrogen stimulus [BP],
metal ion binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_179_4 |
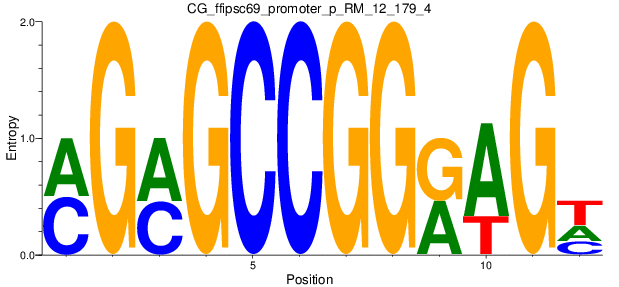 |
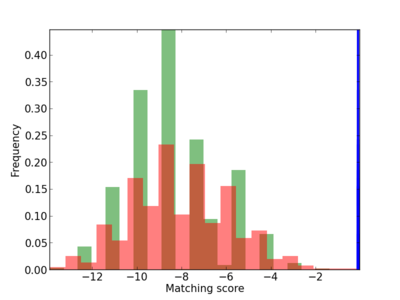 |
6 |
0.8 |
2.48E-04 |
ACSL3,
EPN1,
AP2A2,
PCYOX1 |
positive regulation of phosphatidylcholine biosynthetic process [BP],
chloride-transporting ATPase activity [MF],
negative regulation of epidermal growth factor receptor signaling pathway [BP],
prenylcysteine oxidase activity [MF] |
| CG_ffipsc69_promoter_p_RM_14_10_6 |
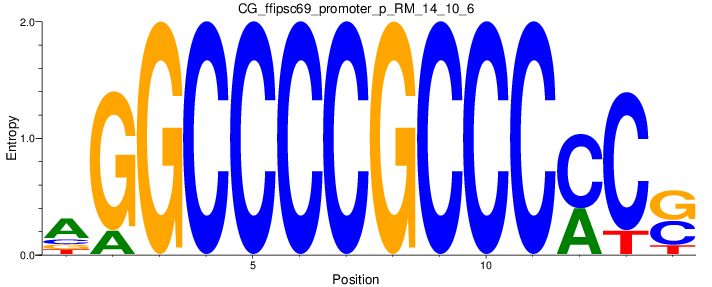 |
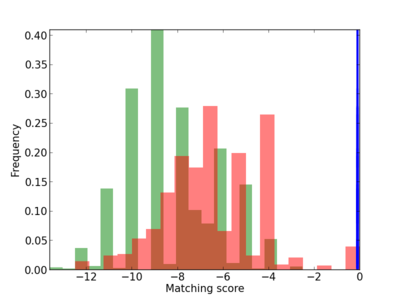 |
7 |
1.0 |
2.38E-04 |
NET1 |
cellular response to ionizing radiation [BP] |
| CG_ffipsc69_promoter_p_RM_12_3_8 |
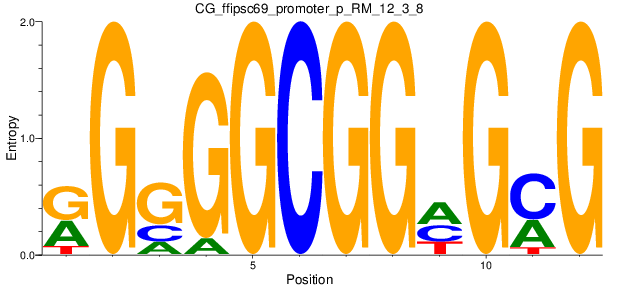 |
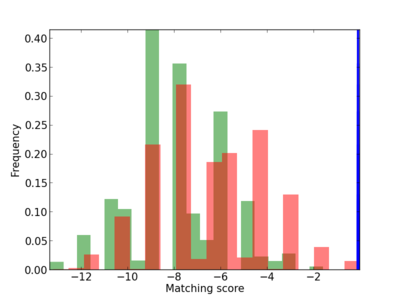 |
4 |
0.6 |
3.73E-04 |
UPF3A,
PPP1R1B,
MT1X,
PCIF1,
NTMT1,
BIRC5,
EAF2,
FGF2,
API5,
PPARD,
ZNF90,
MYNN,
HOXC8,
MLX,
IGFN1,
PHF3,
ZNF771,
GABPB2 |
fibroblast growth factor binding [MF],
nucleus [CC] |
| CG_ffipsc69_promoter_p_RM_12_1_6 |
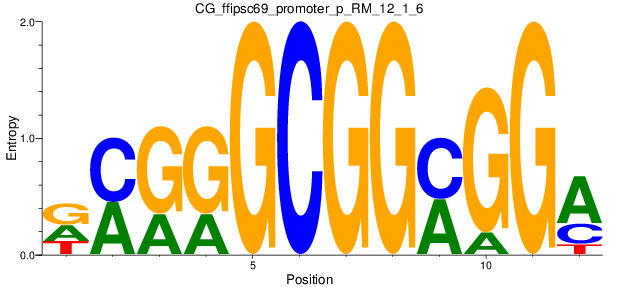 |
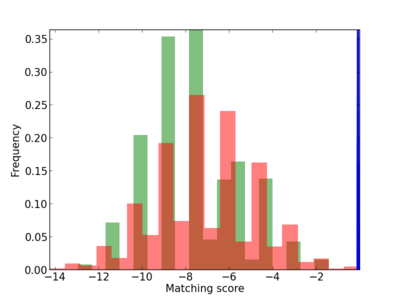 |
5 |
1.0 |
2.32E-04 |
EMG1,
SHROOM2 |
eye pigment granule organization [BP],
rRNA (pseudouridine) methyltransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_14_25_10 |
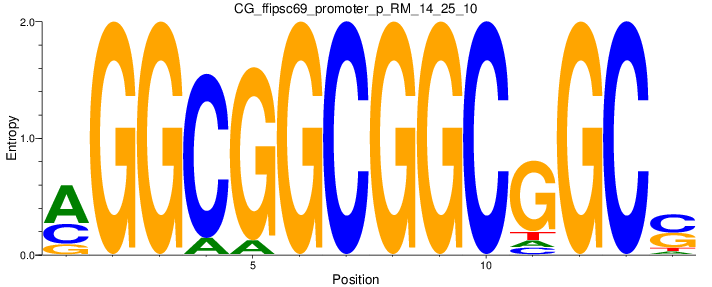 |
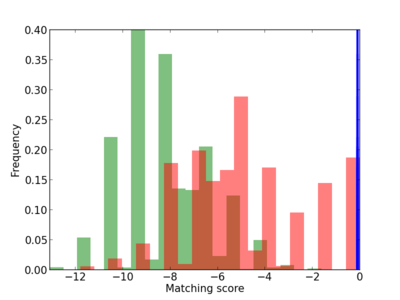 |
13 |
1.0 |
4.06E-04 |
BHLHE22,
AK7,
REST,
SOX8,
ZIC2,
PAX2,
BCL7A,
FZD1 |
ureter development [BP],
brain development [BP],
stem cell differentiation [BP],
negative regulation of transcription, DNA-dependent [BP],
ureter morphogenesis [BP],
metanephric nephron tubule formation [BP] |
| CG_ffipsc69_promoter_p_RM_14_35_2 |
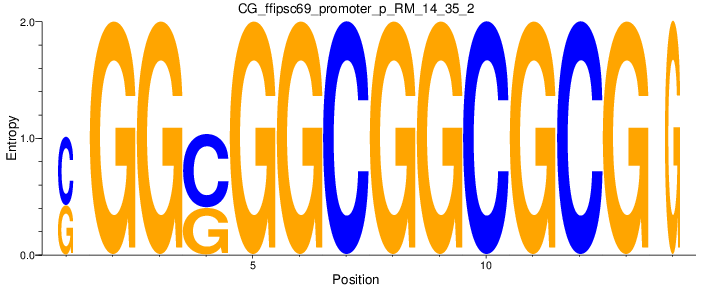 |
 |
3 |
0.4 |
6.84E-04 |
SOX11 |
positive regulation of lens epithelial cell proliferation [BP],
positive regulation of hippo signaling cascade [BP] |
| CG_ffipsc69_promoter_p_RM_14_45_3 |
 |
 |
6 |
0.6 |
2.06E-04 |
TMX1,
UGP2,
PAX2,
HSD3B7 |
cholest-5-ene-3-beta,7-alpha-diol 3-beta-dehydrogenase activity [MF],
UTP:glucose-1-phosphate uridylyltransferase activity [MF],
positive regulation of optic nerve formation [BP],
optic chiasma development [BP],
glycogen biosynthetic process [BP],
arsenate reductase (thioredoxin) activity [MF],
vestibulocochlear nerve formation [BP] |
| CG_ffipsc69_promoter_p_RM_12_72_5 |
 |
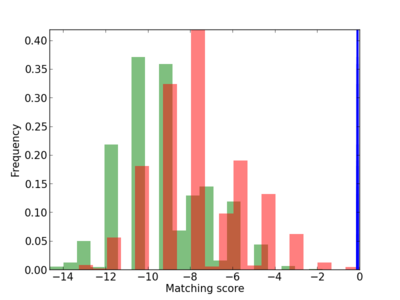 |
4 |
0.8 |
3.25E-04 |
DYNLRB1,
SKIL,
DYNLRB2 |
cytoplasmic dynein complex [CC],
negative regulation of cell differentiation [BP] |
| CG_ffipsc69_promoter_p_RM_12_130_7 |
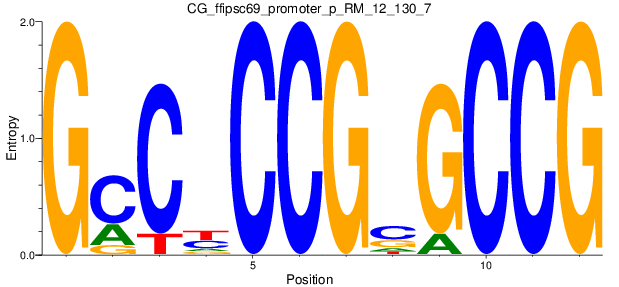 |
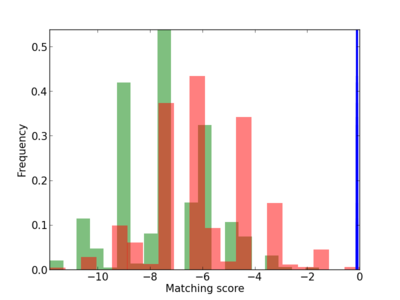 |
3 |
0.4 |
2.98E-04 |
MTRR,
PFAS,
ATP12A,
GMPR2 |
hydrogen:potassium-exchanging ATPase complex [CC],
[methionine synthase] reductase activity [MF],
phosphoribosylformylglycinamidine synthase activity [MF],
purine nucleobase metabolic process [BP] |
| CG_ffipsc69_promoter_p_RM_12_210_3 |
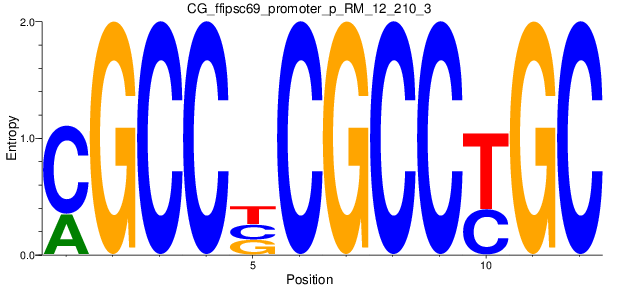 |
 |
2 |
0.4 |
2.87E-04 |
TGIF1,
CYP26A1 |
negative regulation of retinoic acid receptor signaling pathway [BP] |
| CG_ffipsc69_promoter_p_RM_12_262_13 |
 |
 |
5 |
0.8 |
2.44E-04 |
CDC25B,
BLOC1S3,
FOXO3 |
oocyte maturation [BP],
eye development [BP] |
| CG_ffipsc69_promoter_p_RM_12_83_13 |
 |
 |
5 |
1.0 |
2.36E-04 |
PTGFR,
GPT2,
VPS16,
VPS18,
DAND5,
TUBB,
OSTF1,
DYRK2,
NCAPH,
GPT,
SLC48A1,
RELT,
SKI,
SYNGAP1 |
negative regulation of BMP signaling pathway [BP],
HOPS complex [CC],
negative regulation of activin receptor signaling pathway [BP],
L-alanine metabolic process [BP],
L-alanine catabolic process [BP],
cytoplasm [CC],
L-alanine:2-oxoglutarate aminotransferase activity [MF],
endosome membrane [CC] |
| CG_ffipsc69_promoter_p_RM_12_246_10 |
 |
 |
4 |
0.8 |
2.30E-04 |
MAN2A1,
MIF,
VEGFA |
chemoattractant activity [MF],
cell surface receptor signaling pathway [BP],
hydrolase activity, hydrolyzing N-glycosyl compounds [MF] |
| CG_ffipsc69_promoter_p_RM_14_1_17 |
 |
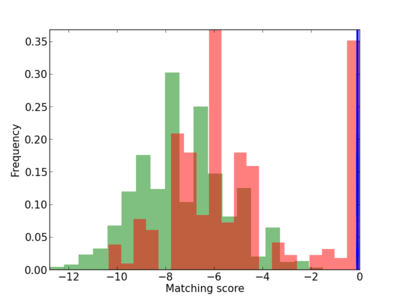 |
7 |
1.0 |
4.72E-04 |
ACVR2A,
EGR1,
GNAS,
BHLHE40,
ARID1A,
PPRC1,
HMGA2,
RING1,
HOXD11,
NELFB,
CUX2,
BMPR2,
MLLT10,
TBX4,
NFATC2IP,
GRHL3,
KCNIP3,
HIVEP1,
PKNOX1,
ARNTL |
RNA polymerase II core promoter proximal region sequence-specific DNA binding transcription factor activity [MF],
activin-activated receptor activity [MF],
positive regulation of osteoblast differentiation [BP],
regulation of transcription, DNA-dependent [BP],
sequence-specific DNA binding transcription factor activity [MF],
transcription, DNA-dependent [BP],
sequence-specific DNA binding RNA polymerase II transcription factor activity [MF],
circadian regulation of gene expression [BP],
nucleic acid binding transcription factor activity [MF],
transcription from RNA polymerase II promoter [BP],
regulation of transcription from RNA polymerase II promoter [BP] |
| CG_ffipsc69_promoter_p_RM_12_216_8 |
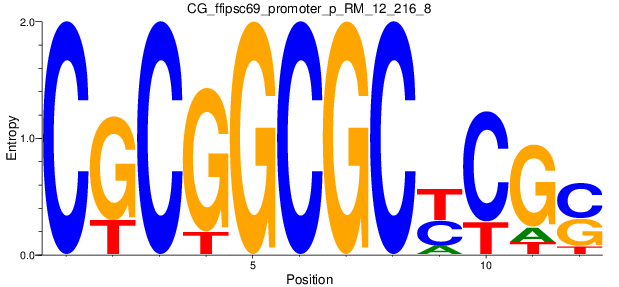 |
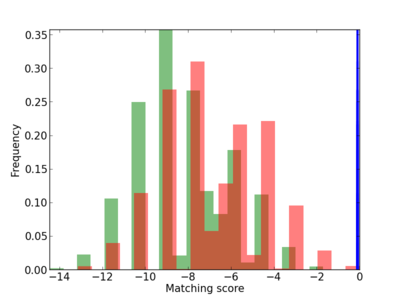 |
4 |
0.6 |
4.25E-04 |
TPST1,
PRKCE |
TRAM-dependent toll-like receptor 4 signaling pathway [BP],
positive regulation of cellular glucuronidation [BP],
ethanol binding [MF],
protein-tyrosine sulfotransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_184_7 |
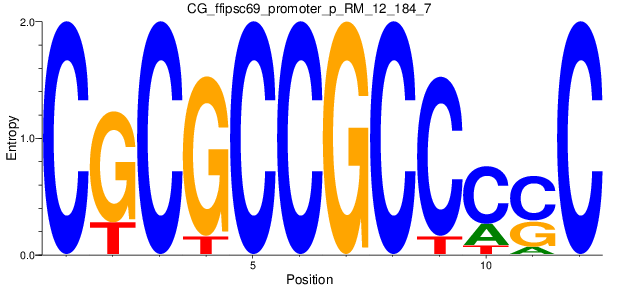 |
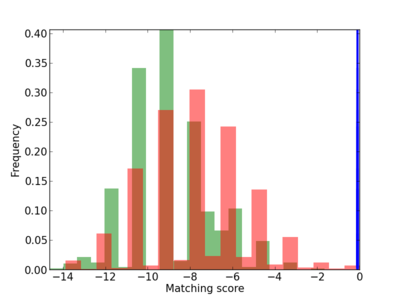 |
4 |
0.8 |
3.74E-04 |
NOG,
FOXL1,
NR2E1,
ZIC1,
HES3,
PAPD5,
EXOSC4,
CHD7 |
regulation of timing of neuron differentiation [BP],
negative regulation of astrocyte differentiation [BP],
brain development [BP],
central nervous system development [BP],
pattern specification process [BP],
histone mRNA catabolic process [BP] |
| CG_ffipsc69_promoter_p_RM_14_47_7 |
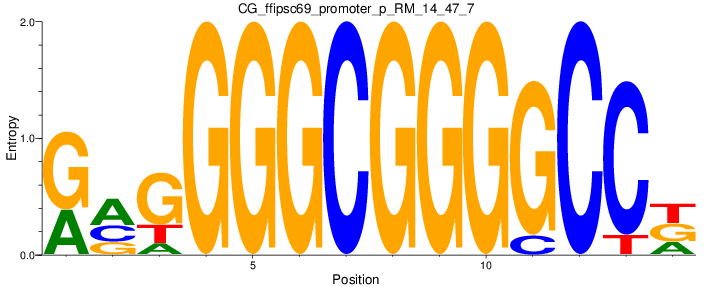 |
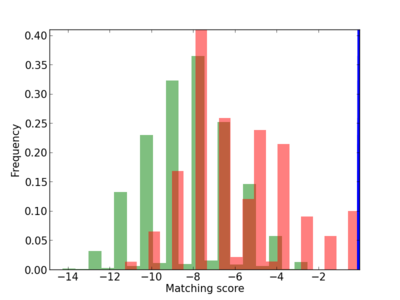 |
9 |
1.0 |
2.94E-04 |
STXBP3,
STXBP2,
GRIN1,
TRPM4 |
neutrophil degranulation [BP],
specific granule [CC],
calcium channel activity [MF],
tertiary granule [CC],
syntaxin-1 binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_175_6 |
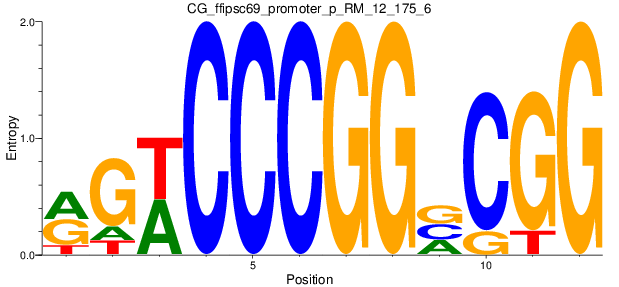 |
 |
2 |
0.4 |
1.77E-04 |
CTCF,
KAT6B,
ING5 |
protein acetylation [BP],
MOZ/MORF histone acetyltransferase complex [CC],
histone H3 acetylation [BP],
chromatin modification [BP],
histone acetylation [BP] |
| CG_ffipsc69_promoter_p_RM_12_112_10 |
 |
 |
5 |
0.8 |
1.23E-04 |
SPEG |
muscle organ development [BP],
muscle cell differentiation [BP] |
| CG_ffipsc69_promoter_p_RM_12_59_6 |
 |
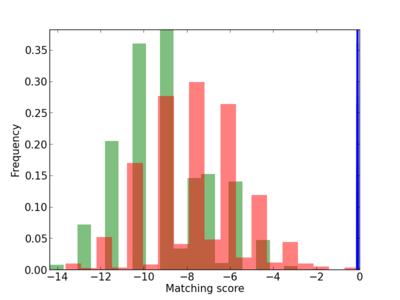 |
3 |
0.6 |
3.48E-04 |
TRAPPC10,
SPINT1 |
placenta blood vessel development [BP],
sodium ion transmembrane transporter activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_56_4 |
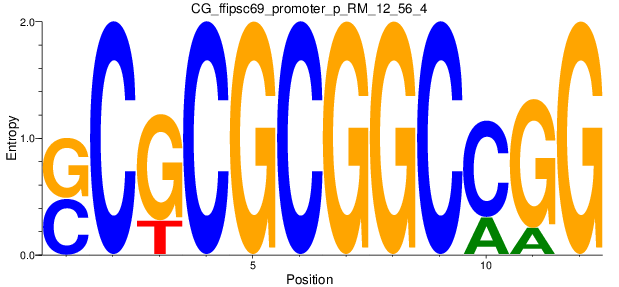 |
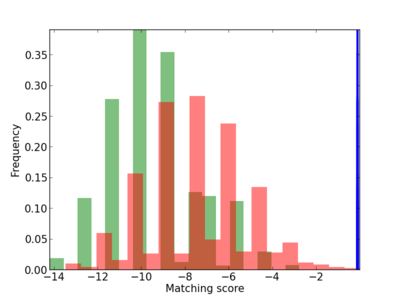 |
5 |
1.0 |
2.36E-04 |
TBK1,
IRF5,
ADAMTSL4,
VWC2 |
positive regulation of interferon-alpha production [BP],
positive regulation of interferon-beta production [BP],
interstitial matrix [CC] |
| CG_ffipsc69_promoter_p_RM_12_286_5 |
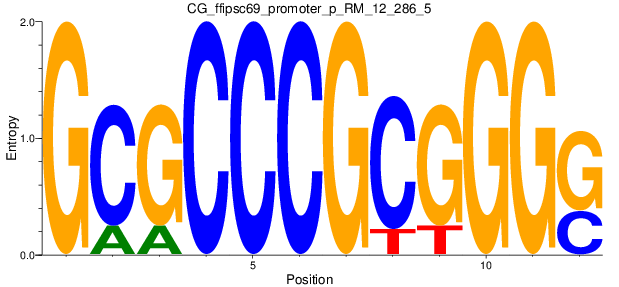 |
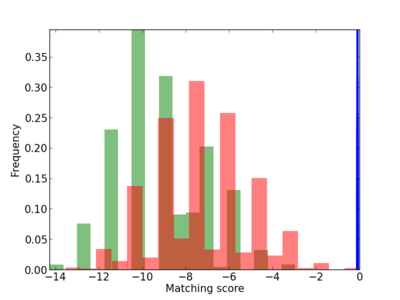 |
2 |
0.4 |
4.97E-04 |
TARBP2 |
miRNA loading onto RISC involved in gene silencing by miRNA [BP] |
| CG_ffipsc69_promoter_p_RM_12_156_10 |
 |
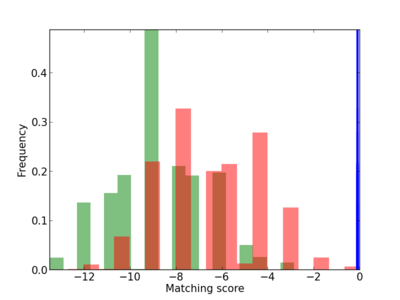 |
3 |
0.6 |
3.78E-04 |
ZDHHC5,
SYNGR3,
HIF1A,
DAG1,
GATA6,
ACVRL1 |
regulation of transforming growth factor beta2 production [BP],
SH2 domain binding [MF],
intestinal epithelial cell differentiation [BP],
cellular response to BMP stimulus [BP],
retina vasculature development in camera-type eye [BP],
palmitoyltransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_182_6 |
 |
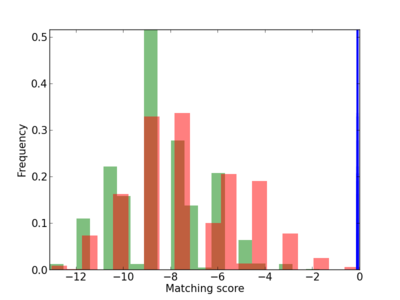 |
7 |
1.0 |
3.82E-04 |
PNP,
CYR61,
CRTAP,
SRD5A3 |
NAD biosynthesis via nicotinamide riboside salvage pathway [BP],
polyprenol catabolic process [BP],
purine-nucleoside phosphorylase activity [MF],
inosine catabolic process [BP],
intussusceptive angiogenesis [BP],
nicotinamide riboside catabolic process [BP],
peptidyl-proline hydroxylation to 3-hydroxy-L-proline [BP] |
| CG_ffipsc69_promoter_p_RM_14_24_8 |
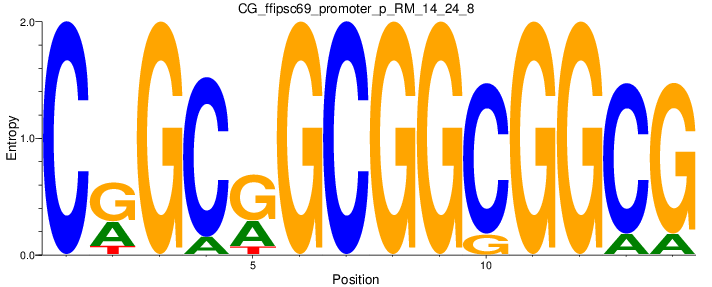 |
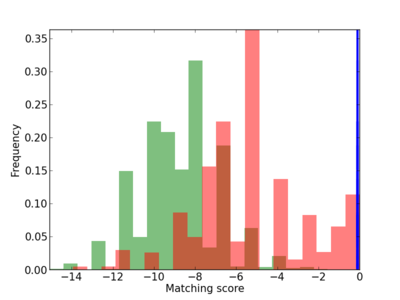 |
13 |
1.0 |
3.88E-04 |
PHF17,
SUPT6H,
TSPYL2,
CDC73,
PPP5C,
FZD9,
ARF6,
SOX8,
NAA15,
ZIC2,
RTF1,
SALL3,
FUBP3,
ZMIZ1,
CNOT6,
ZFY |
filopodium membrane [CC],
regulation of DNA-dependent transcription, elongation [BP],
regulation of transcription, DNA-dependent [BP],
transcription, DNA-dependent [BP],
Cdc73/Paf1 complex [CC],
positive regulation of transcription elongation from RNA polymerase II promoter [BP],
RNA metabolic process [BP],
endodermal cell fate commitment [BP] |
| CG_ffipsc69_promoter_p_RM_12_250_13 |
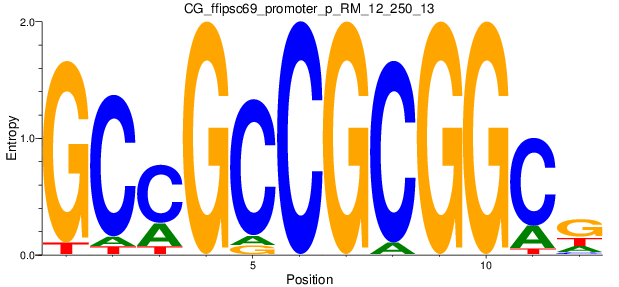 |
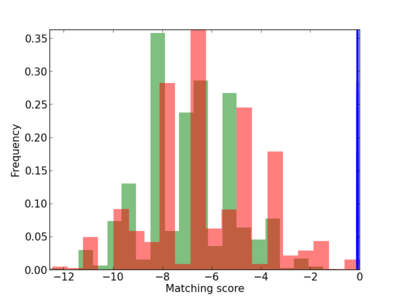 |
5 |
1.0 |
3.96E-04 |
EGR1,
SOX9 |
cellular response to heparin [BP] |
| CG_ffipsc69_promoter_p_RM_12_71_9 |
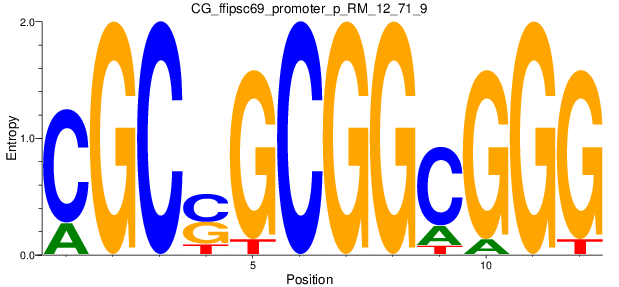 |
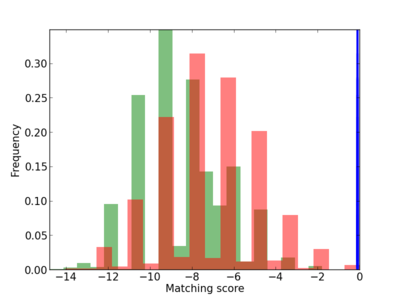 |
4 |
0.8 |
2.90E-04 |
HSPA6,
COPS4,
GPS1 |
cullin deneddylation [BP],
signalosome [CC] |
| CG_ffipsc69_promoter_p_RM_12_103_7 |
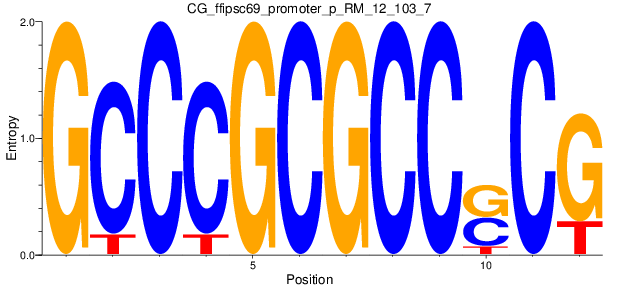 |
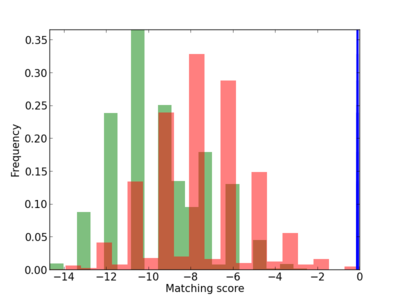 |
3 |
0.6 |
3.12E-04 |
NARF,
PLCB1,
WBP2NL |
lamin binding [MF],
egg activation [BP] |
| CG_ffipsc69_promoter_p_RM_12_45_5 |
 |
 |
4 |
0.6 |
2.31E-04 |
SOX4,
NAA15 |
N-terminal protein amino acid acetylation [BP],
core promoter sequence-specific DNA binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_152_8 |
 |
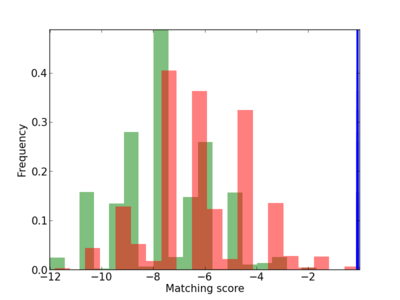 |
5 |
0.8 |
3.10E-04 |
HEY2 |
negative regulation of cardiac vascular smooth muscle cell differentiation [BP] |
| CG_ffipsc69_promoter_p_RM_14_51_3 |
 |
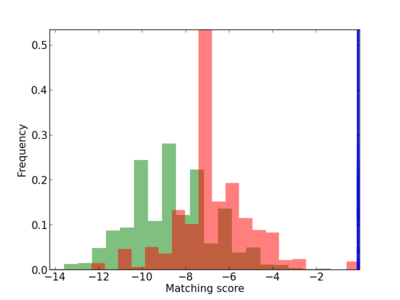 |
4 |
0.6 |
1.30E-04 |
NQO2,
SGPL1 |
dihydronicotinamide riboside quinone reductase activity [MF],
NADPH dehydrogenase (quinone) activity [MF],
sphinganine-1-phosphate aldolase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_226_4 |
 |
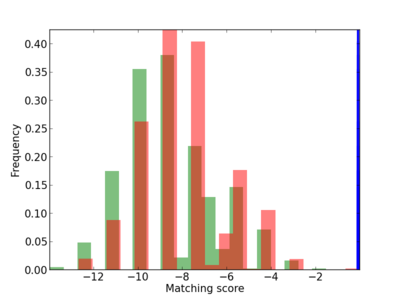 |
7 |
0.8 |
2.76E-04 |
TNFSF11,
RGL1 |
Ral guanyl-nucleotide exchange factor activity [MF],
positive regulation of fever generation by positive regulation of prostaglandin secretion [BP],
TNFSF11-mediated signaling pathway [BP],
positive regulation of ERK1 and ERK2 cascade via TNFSF11-mediated signaling [BP],
positive regulation of corticotropin-releasing hormone secretion [BP] |
| CG_ffipsc69_promoter_p_RM_12_278_5 |
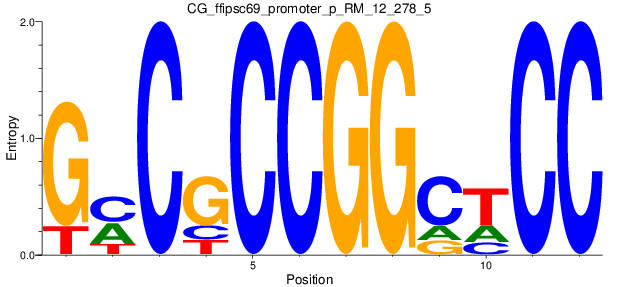 |
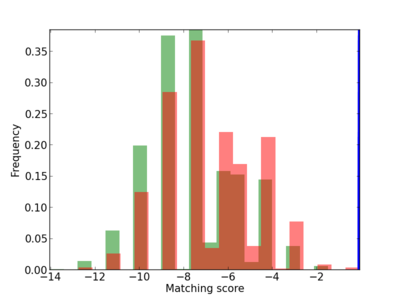 |
2 |
0.4 |
2.09E-04 |
NMT1 |
N-terminal protein myristoylation [BP],
glycylpeptide N-tetradecanoyltransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_145_6 |
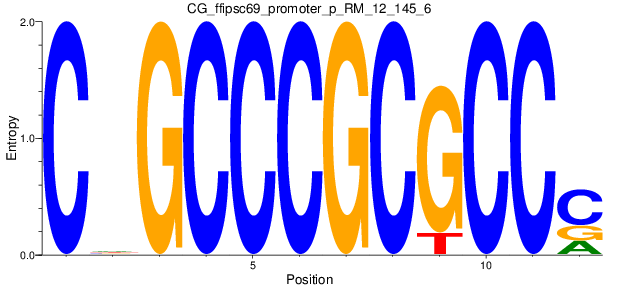 |
 |
8 |
1.0 |
2.86E-04 |
MIXL1 |
cell migration involved in gastrulation [BP] |
| CG_ffipsc69_promoter_p_RM_12_276_2 |
 |
 |
1 |
0.2 |
5.17E-04 |
PIN1,
YY1,
WNK2 |
negative regulation of ERK1 and ERK2 cascade [BP],
response to prostaglandin F stimulus [BP] |
| CG_ffipsc69_promoter_p_RM_12_193_7 |
 |
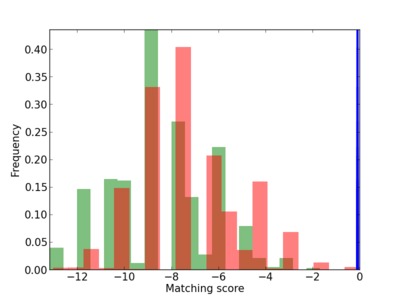 |
4 |
0.8 |
3.81E-04 |
ID2,
DLL3 |
positive regulation of transcription involved in G1/S phase of mitotic cell cycle [BP],
negative regulation of neurogenesis [BP] |
| CG_ffipsc69_promoter_p_RM_12_180_4 |
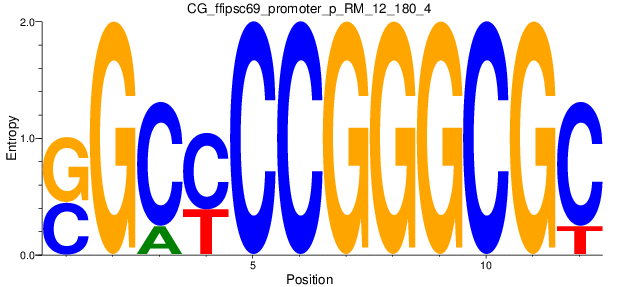 |
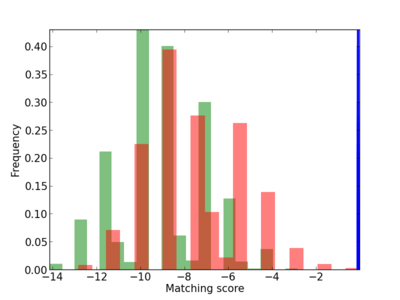 |
4 |
0.6 |
3.33E-04 |
CNTN2,
SMARCA2,
SLC35A1,
BTD,
CHST11 |
biotinidase activity [MF],
CMP-N-acetylneuraminate transmembrane transporter activity [MF],
positive regulation of adenosine receptor signaling pathway [BP],
aortic smooth muscle cell differentiation [BP],
polysaccharide localization [BP],
CMP-N-acetylneuraminate transport [BP],
establishment of protein localization to juxtaparanode region of axon [BP] |
| CG_ffipsc69_promoter_p_RM_12_46_5 |
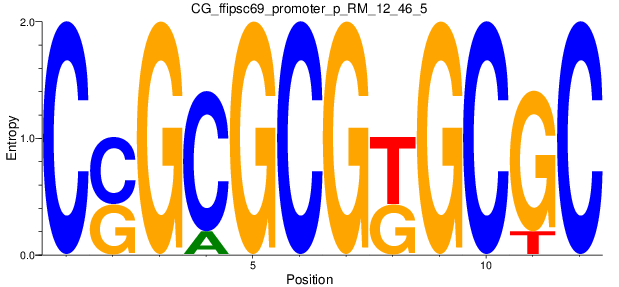 |
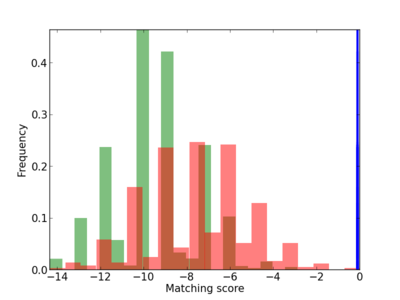 |
3 |
0.6 |
4.11E-04 |
MCM4,
FOXD3,
NF1,
TRIM32,
POLA2 |
DNA replication initiation [BP],
sympathetic nervous system development [BP],
negative regulation of fibroblast proliferation [BP] |
| CG_ffipsc69_promoter_p_RM_14_40_1 |
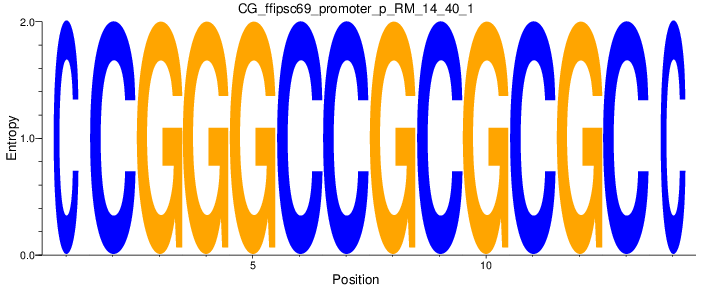 |
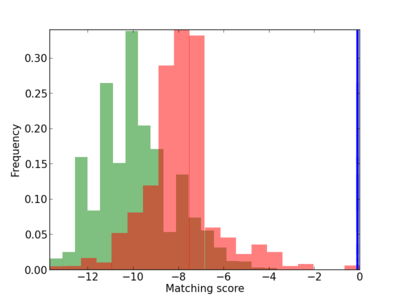 |
3 |
0.6 |
3.19E-04 |
MCCC2,
DAG1 |
methylcrotonoyl-CoA carboxylase activity [MF],
nerve maturation [BP] |
| CG_ffipsc69_promoter_p_RM_12_251_7 |
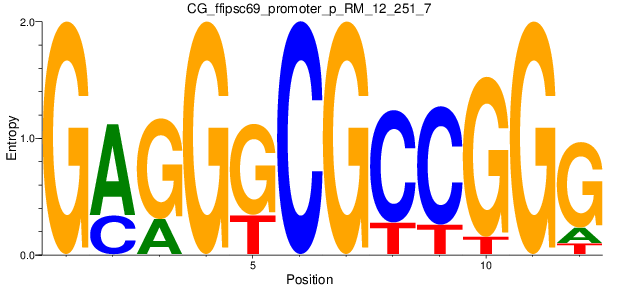 |
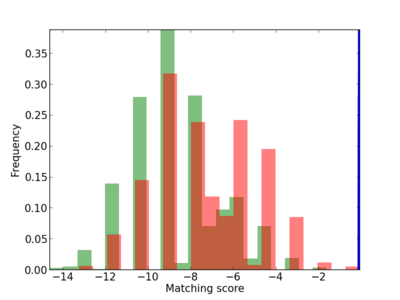 |
3 |
0.6 |
2.21E-04 |
HES1 |
somatic stem cell maintenance [BP] |
| CG_ffipsc69_promoter_p_RM_12_163_6 |
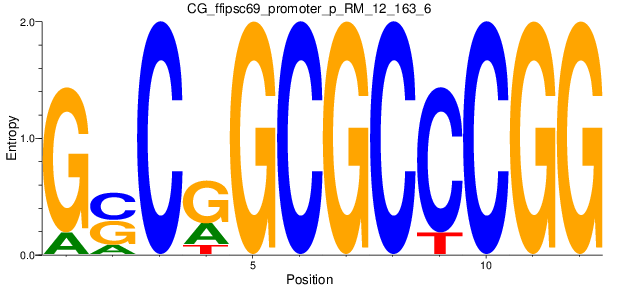 |
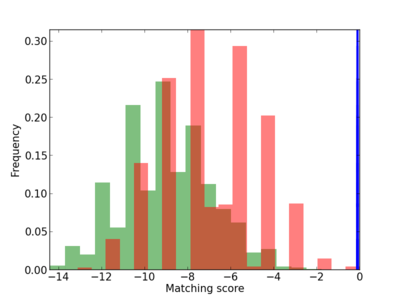 |
3 |
0.6 |
2.40E-04 |
RUNX2,
JAK2,
HPN,
SALL3,
GNPAT,
CR1 |
complement component C4b binding [MF],
complement component C3b receptor activity [MF],
positive regulation of serine-type endopeptidase activity [BP],
negative regulation of smoothened signaling pathway [BP],
negative regulation of complement activation, alternative pathway [BP],
mineralocorticoid receptor signaling pathway [BP],
glycerone-phosphate O-acyltransferase activity [MF],
histone kinase activity (H3-Y41 specific) [MF],
complement component C4b receptor activity [MF],
histone H3-Y41 phosphorylation [BP],
complement component C3b binding [MF],
basement membrane disassembly [BP],
activation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway [BP],
negative regulation of serine-type endopeptidase activity [BP],
negative regulation of complement activation, classical pathway [BP] |
| CG_ffipsc69_promoter_p_RM_12_150_5 |
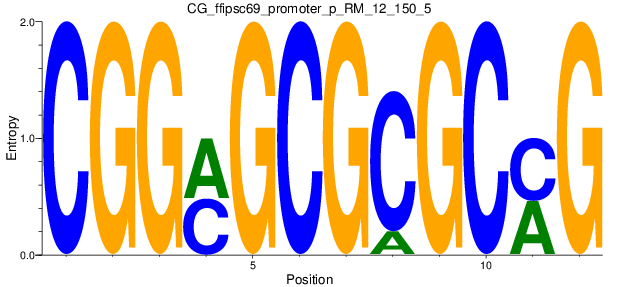 |
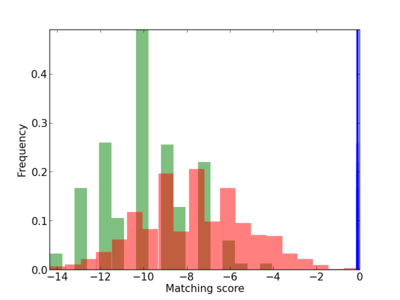 |
3 |
0.6 |
2.58E-04 |
SYNDIG1 |
glutamate receptor binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_43_5 |
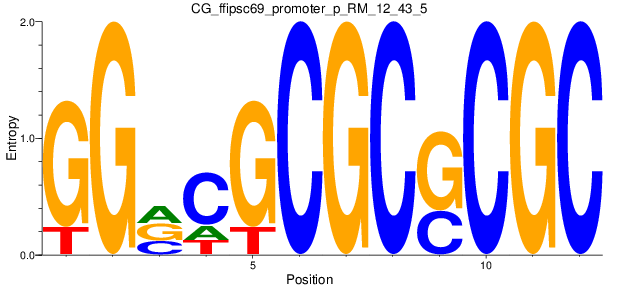 |
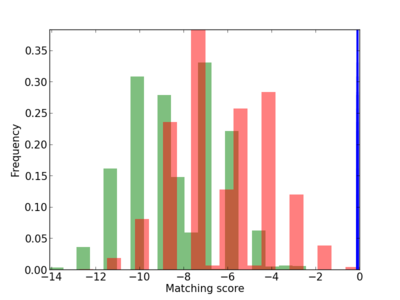 |
5 |
0.8 |
2.54E-04 |
ZNF865,
ZFP1,
TNFAIP3,
PRKAA2,
RNF4 |
protein K48-linked ubiquitination [BP],
regulation of transcription, DNA-dependent [BP] |
| CG_ffipsc69_promoter_p_RM_12_287_7 |
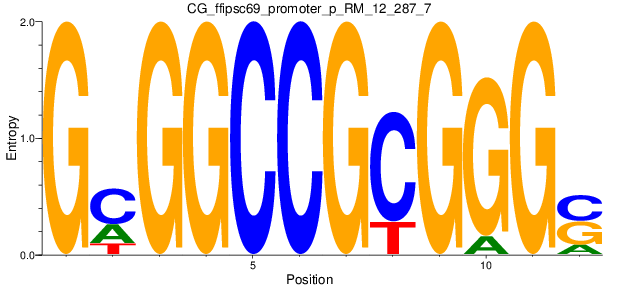 |
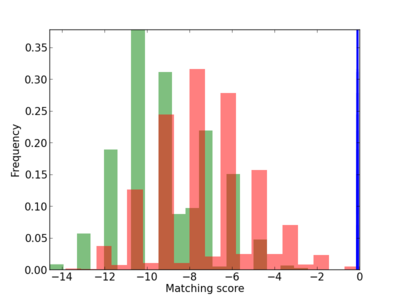 |
6 |
0.8 |
2.38E-04 |
PTF1A |
retinoic acid receptor signaling pathway [BP] |
| CG_ffipsc69_promoter_p_RM_12_267_6 |
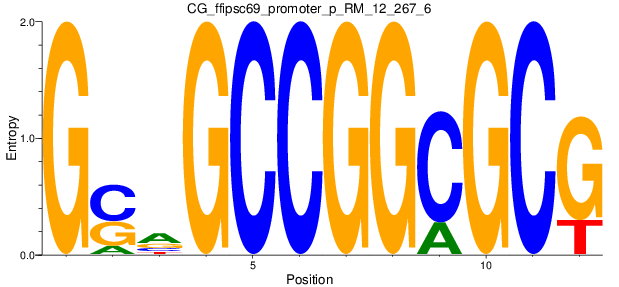 |
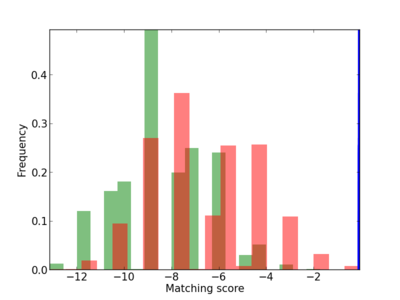 |
4 |
0.8 |
2.49E-04 |
LHX9,
SMAD6,
ILK,
MSMO1,
POU3F3 |
ureteric bud development [BP],
gonad morphogenesis [BP],
metanephric macula densa development [BP],
neuron projection morphogenesis [BP],
C-4 methylsterol oxidase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_268_10 |
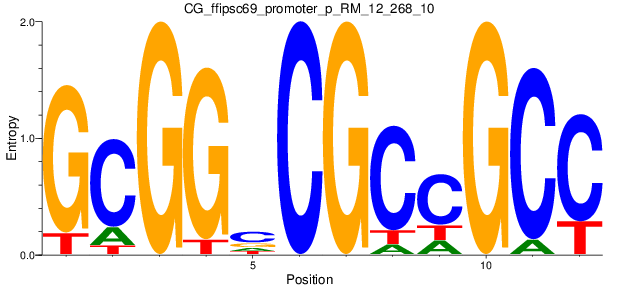 |
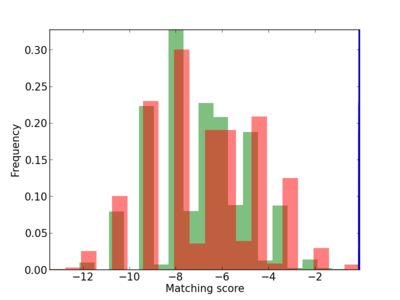 |
6 |
1.0 |
7.02E-04 |
PCSK9,
NTRK1,
SOX8 |
negative regulation of receptor recycling [BP],
Sertoli cell development [BP],
retinal rod cell differentiation [BP],
neurotrophin p75 receptor binding [MF],
male gonad development [BP],
nerve growth factor receptor activity [MF],
very-low-density lipoprotein particle binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_116_5 |
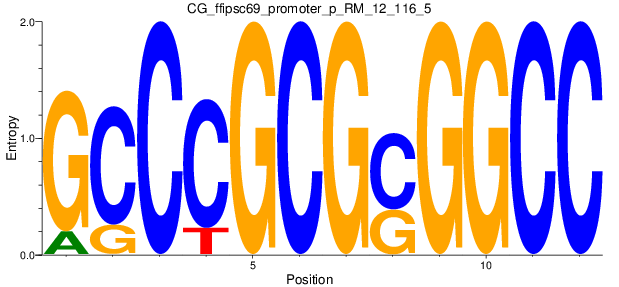 |
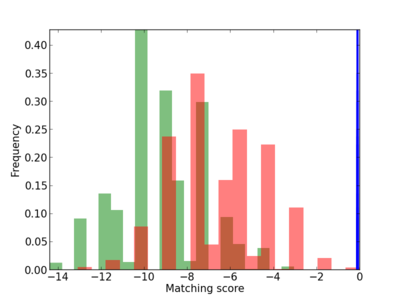 |
3 |
0.6 |
1.92E-04 |
MGST1,
ARFGEF2,
EGR1,
GAA,
ICT1 |
symmetric synapse [CC],
diaphragm contraction [BP],
sucrose metabolic process [BP],
translation release factor activity, codon nonspecific [MF],
cellular response to lipid hydroperoxide [BP],
maltose metabolic process [BP],
positive regulation of glomerular metanephric mesangial cell proliferation [BP],
vacuolar sequestering [BP],
mitochondrial translational termination [BP] |
| CG_ffipsc69_promoter_p_RM_12_31_3 |
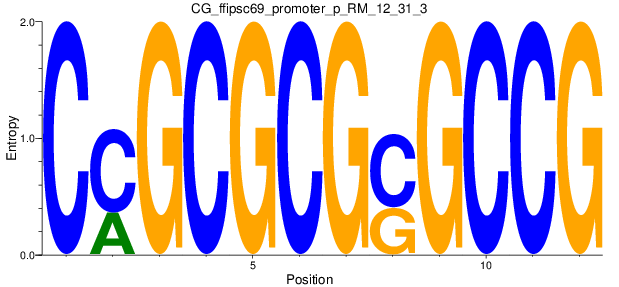 |
 |
2 |
0.4 |
2.34E-04 |
TM7SF2,
PRKAA2,
SLC27A1 |
long-chain fatty acid transport [BP],
cholesterol biosynthetic process [BP] |
| CG_ffipsc69_promoter_p_RM_12_91_6 |
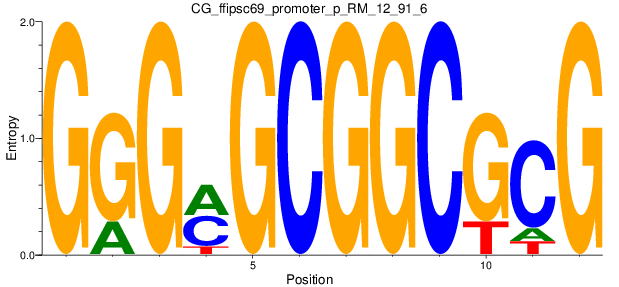 |
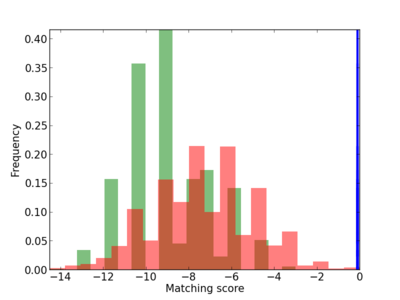 |
3 |
0.6 |
3.88E-04 |
SSH3 |
regulation of lamellipodium assembly [BP] |
| CG_ffipsc69_promoter_p_RM_12_73_8 |
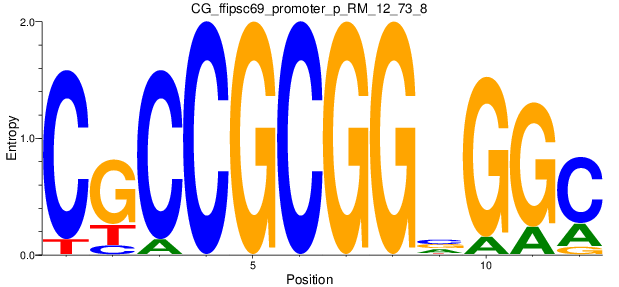 |
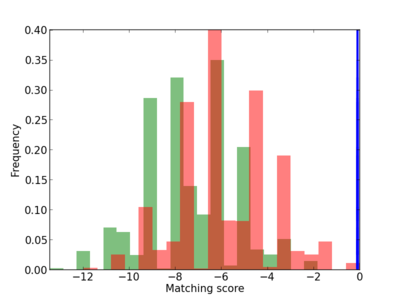 |
6 |
1.0 |
4.17E-04 |
CEBPG,
EGR1,
ZBTB8A,
ZIC2,
HIST1H2AH,
MLL4,
ELP2,
ZBTB1 |
DNA binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_197_2 |
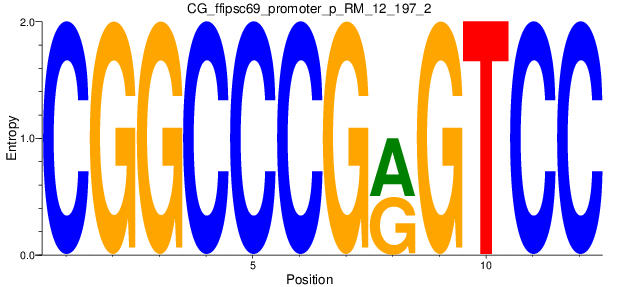 |
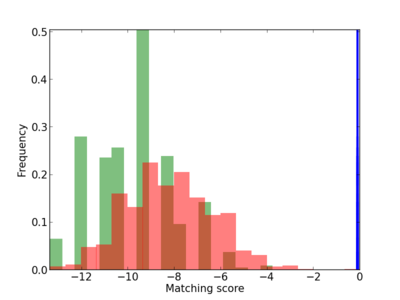 |
2 |
0.4 |
2.10E-04 |
PTDSS2,
ALDH2,
ALDH3B1 |
ethanol catabolic process [BP],
CDP-diacylglycerol-serine O-phosphatidyltransferase activity [MF],
aldehyde dehydrogenase [NAD(P)+] activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_54_12 |
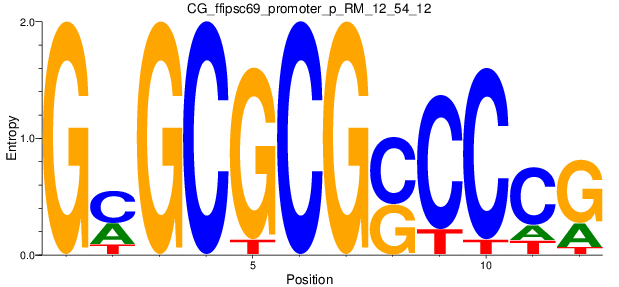 |
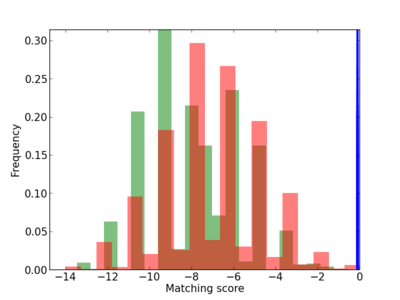 |
5 |
0.8 |
2.99E-04 |
ZCCHC2 |
cell communication [BP] |
| CG_ffipsc69_promoter_p_RM_12_232_12 |
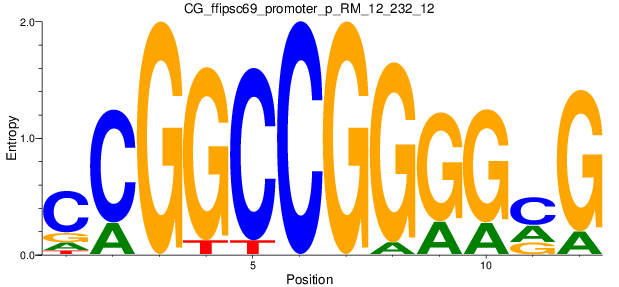 |
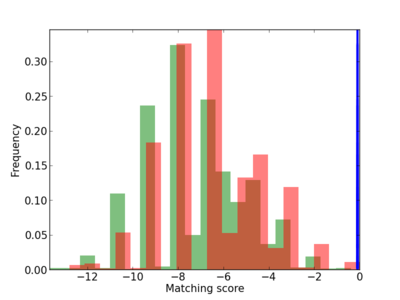 |
3 |
0.6 |
2.15E-04 |
DNAJB2,
PTEN |
postsynaptic density assembly [BP],
negative regulation of cyclin-dependent protein kinase activity involved in G1/S [BP],
inositol-1,3,4,5-tetrakisphosphate 3-phosphatase activity [MF],
negative regulation of ribosome biogenesis [BP],
phosphatidylinositol-3,4-bisphosphate 3-phosphatase activity [MF],
positive regulation of proteasomal ubiquitin-dependent protein catabolic process [BP],
negative regulation of synaptic vesicle clustering [BP] |
| CG_ffipsc69_promoter_p_RM_12_27_3 |
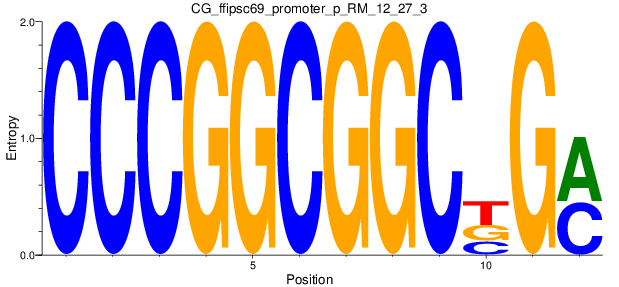 |
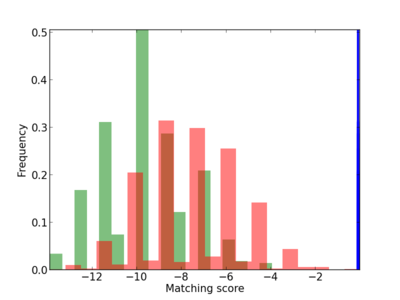 |
4 |
0.6 |
3.56E-04 |
MAPK14,
CACNA1B,
DYNLRB1,
CACNA1H |
voltage-gated calcium channel activity [MF],
voltage-gated calcium channel complex [CC],
transport [BP],
muscle cell differentiation [BP] |
| CG_ffipsc69_promoter_p_RM_14_39_1 |
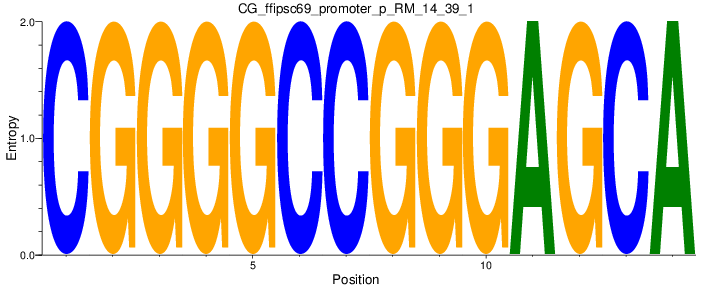 |
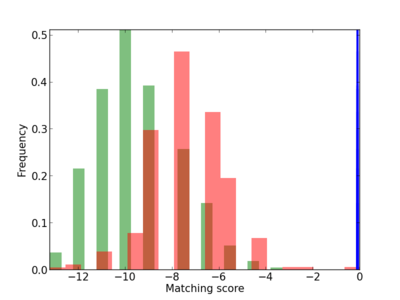 |
5 |
0.8 |
1.80E-04 |
NF2,
MYO3A |
filopodium [CC],
plus-end directed microfilament motor activity [MF],
negative regulation of tyrosine phosphorylation of Stat5 protein [BP] |
| CG_ffipsc69_promoter_p_RM_12_140_5 |
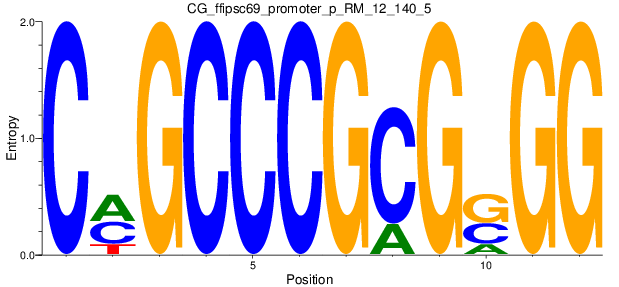 |
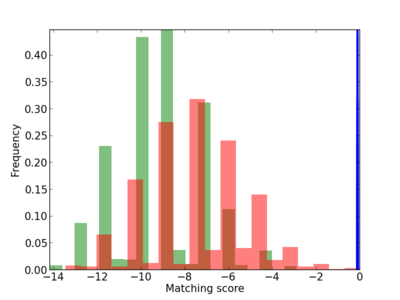 |
5 |
1.0 |
1.80E-04 |
GATA4 |
lung lobe formation [BP] |
| CG_ffipsc69_promoter_p_RM_12_110_9 |
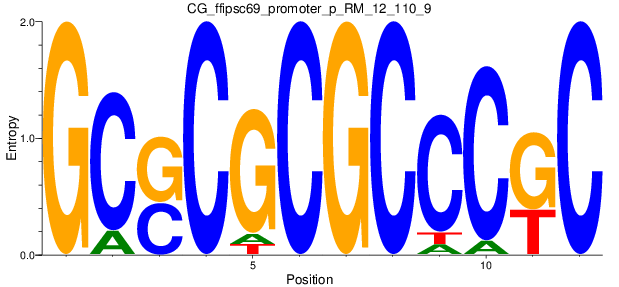 |
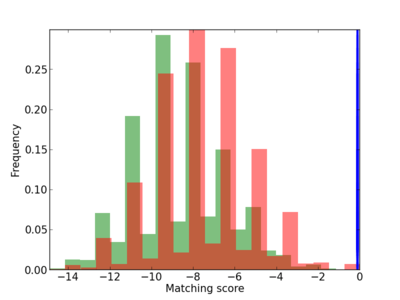 |
4 |
0.8 |
3.73E-04 |
CEBPB,
LTBP4,
SAP30,
UVSSA,
CHD7,
USP1,
STK11,
ENPP1,
MARCH5,
ARHGAP33,
COL1A2,
HTRA2,
UBXN4,
KAT6B,
MAVS,
CLDN4,
RGS9,
FOXC1,
PTPRG,
AMOTL1 |
protein binding [MF],
identical protein binding [MF],
growth hormone secretion [BP],
protein complex binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_21_12 |
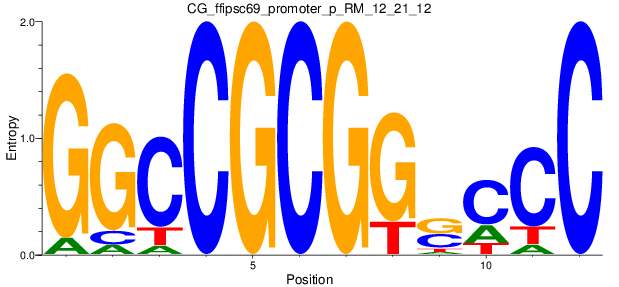 |
 |
4 |
0.8 |
2.32E-04 |
FEM1B,
HHEX |
hepatoblast differentiation [BP],
hepatic duct development [BP],
regulation of DNA damage checkpoint [BP] |
| CG_ffipsc69_promoter_p_RM_12_104_4 |
 |
 |
5 |
0.8 |
3.63E-04 |
PKD2,
CACNA1H |
voltage-gated calcium channel activity [MF],
calcium ion import [BP] |
| CG_ffipsc69_promoter_p_RM_12_263_4 |
 |
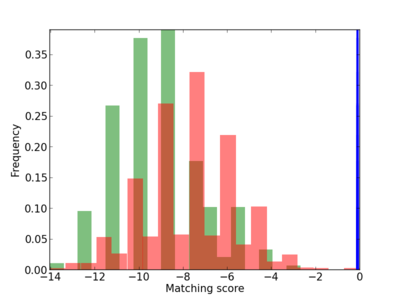 |
4 |
0.8 |
1.26E-04 |
TRNT1,
RARS |
tRNA binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_249_14 |
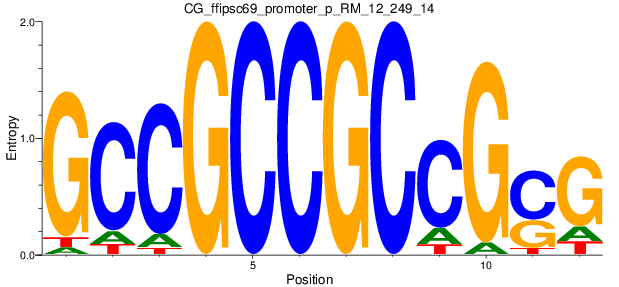 |
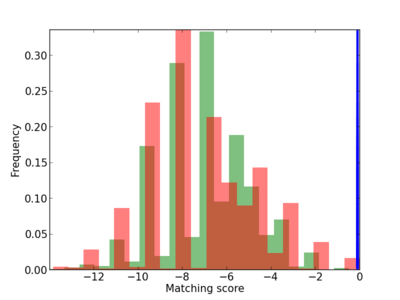 |
4 |
0.8 |
3.61E-04 |
ADORA2A,
FZD2,
WDR36,
BHLHE40,
USP1,
CAPZA2,
SCNN1D,
SPTBN1,
VASP,
NRTN,
ARNTL |
neuron projection development [BP],
neuron projection morphogenesis [BP],
monoubiquitinated protein deubiquitination [BP],
neuron projection membrane [CC],
response to stimulus [BP],
cortical cytoskeleton [CC],
circadian regulation of gene expression [BP],
actin polymerization or depolymerization [BP],
axolemma [CC] |
| CG_ffipsc69_promoter_p_RM_14_22_2 |
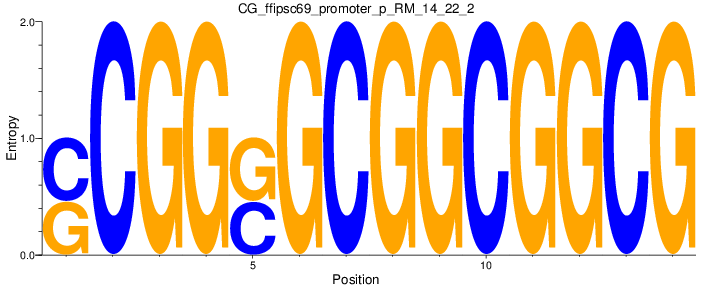 |
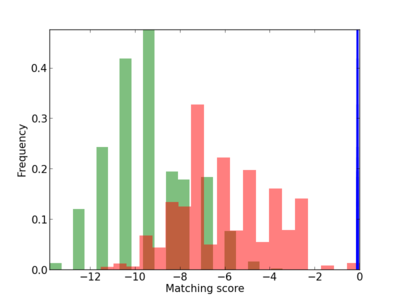 |
3 |
0.4 |
5.66E-04 |
RPS6KB1,
FMN2,
NANOS3 |
oogenesis [BP],
meiotic chromosome movement towards spindle pole [BP],
response to electrical stimulus involved in regulation of muscle adaptation [BP] |
| CG_ffipsc69_promoter_p_RM_12_228_10 |
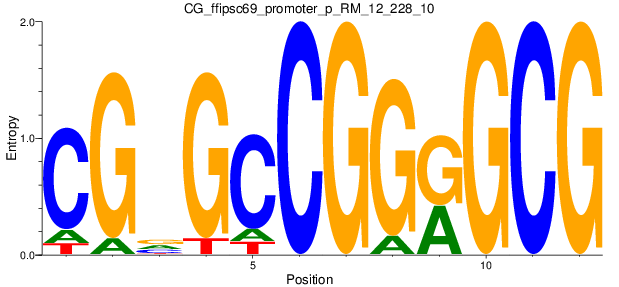 |
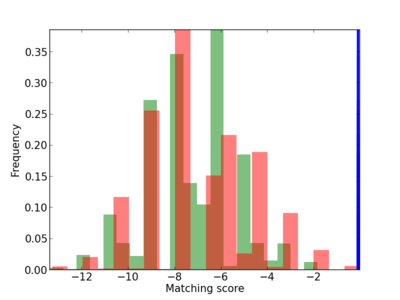 |
3 |
0.6 |
3.43E-04 |
BICD1 |
regulation of proteinase activated receptor activity [BP],
negative regulation of phospholipase C activity [BP],
proteinase activated receptor binding [MF],
negative regulation of phospholipase C-activating G-protein coupled receptor signaling pathway [BP],
host cell viral assembly compartment [CC] |
| CG_ffipsc69_promoter_p_RM_12_288_4 |
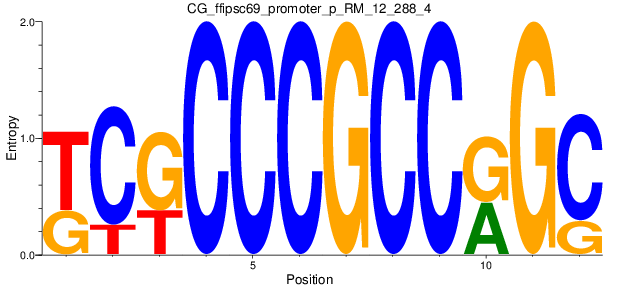 |
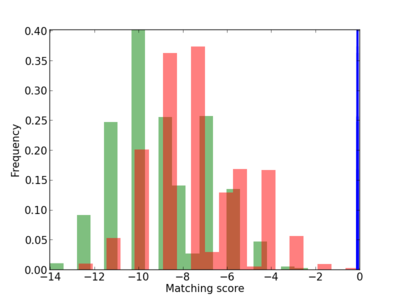 |
3 |
0.4 |
2.51E-04 |
PLOD1,
CACNA1C |
hydroxylysine biosynthetic process [BP],
caveolar macromolecular signaling complex [CC],
calcium-mediated signaling using extracellular calcium source [BP] |
| CG_ffipsc69_promoter_p_RM_14_33_4 |
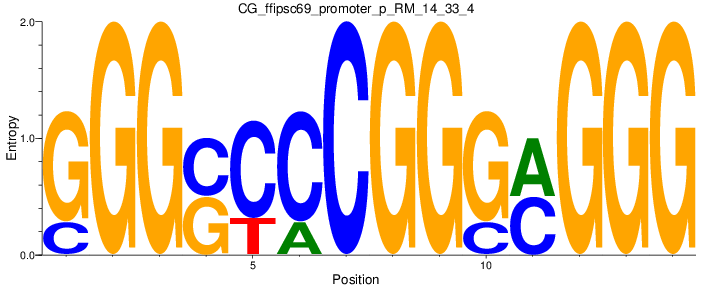 |
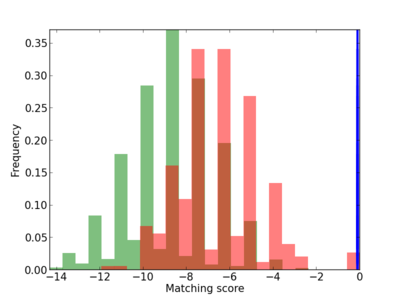 |
4 |
0.4 |
1.57E-04 |
NR2F2,
FZD7 |
radial pattern formation [BP],
negative regulation of ectodermal cell fate specification [BP] |
| CG_ffipsc69_promoter_p_RM_12_90_13 |
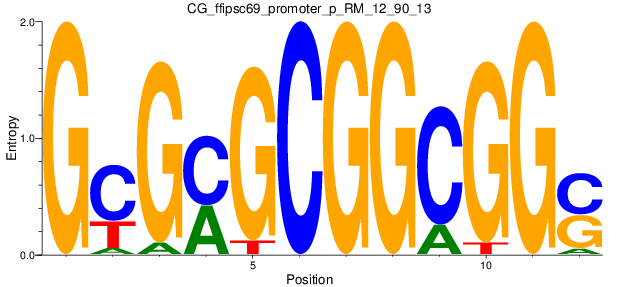 |
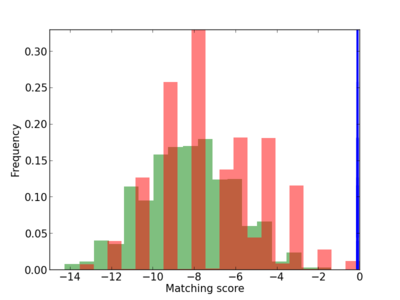 |
6 |
0.8 |
3.62E-04 |
BMPR1B,
ACVRL1,
IQGAP2,
GPS1 |
transmembrane receptor protein serine/threonine kinase activity [MF],
transforming growth factor beta-activated receptor activity [MF],
GTPase inhibitor activity [MF],
transforming growth factor beta receptor activity, type I [MF] |
| CG_ffipsc69_promoter_p_RM_12_160_5 |
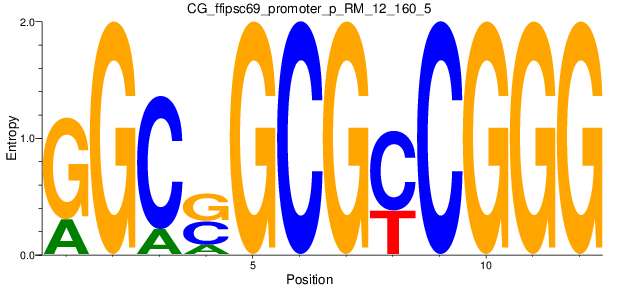 |
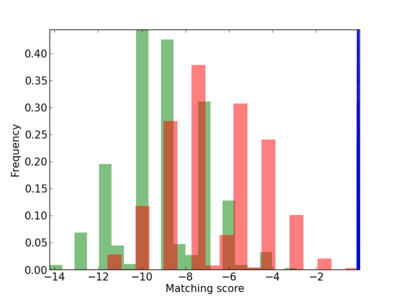 |
6 |
1.0 |
3.49E-04 |
GLIS2,
FLI1,
PPARD,
FOXF2,
BCL6B,
NFIX,
FGFR3,
PDE8B,
CIDEA,
ZBTB10 |
negative regulation of smoothened signaling pathway [BP],
negative regulation of transcription from RNA polymerase II promoter [BP],
regulation of transcription, DNA-dependent [BP],
transcription, DNA-dependent [BP],
negative regulation of transcription, DNA-dependent [BP],
gene expression [BP],
transcription from RNA polymerase II promoter [BP],
regulation of transcription from RNA polymerase II promoter [BP] |
| CG_ffipsc69_promoter_p_RM_14_44_1 |
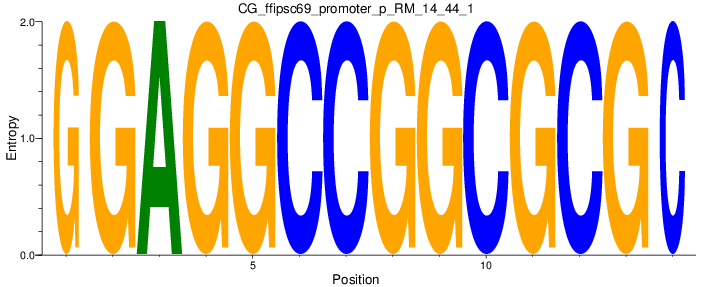 |
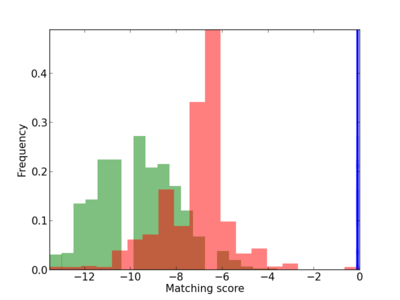 |
2 |
0.4 |
1.93E-04 |
HINFP,
NPM1 |
transcription coactivator activity [MF],
ribosomal large subunit binding [MF],
regulation of endodeoxyribonuclease activity [BP],
regulation of endoribonuclease activity [BP],
histone binding [MF],
negative regulation of centrosome duplication [BP] |
| CG_ffipsc69_promoter_p_RM_12_294_2 |
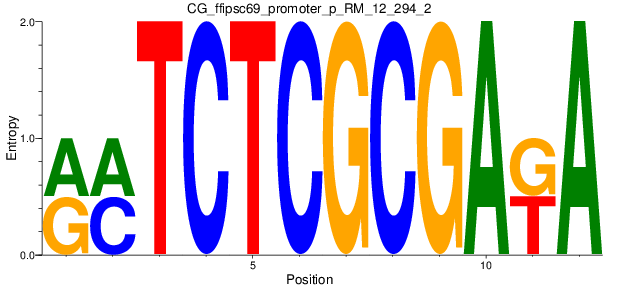 |
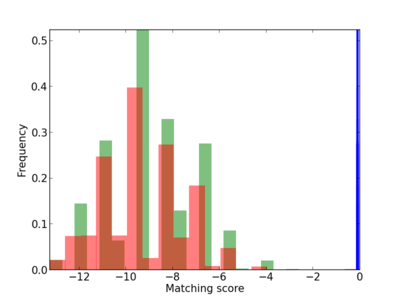 |
4 |
0.8 |
4.24E-04 |
NUDT1,
MOCS3 |
8-oxo-7,8-dihydrodeoxyguanosine triphosphate pyrophosphatase activity [MF],
URM1 activating enzyme activity [MF],
enzyme active site formation via L-cysteine persulfide [BP],
8-oxo-7,8-dihydroguanosine triphosphate pyrophosphatase activity [MF],
ATP diphosphatase activity [MF] |
| CG_ffipsc69_promoter_p_RM_14_8_2 |
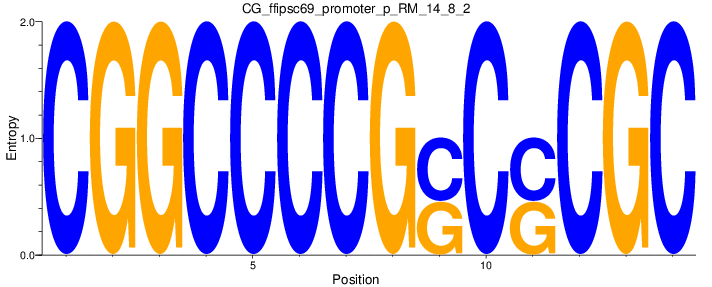 |
 |
4 |
0.8 |
4.29E-04 |
TEAD4 |
hippo signaling cascade [BP] |
| CG_ffipsc69_promoter_p_RM_12_289_5 |
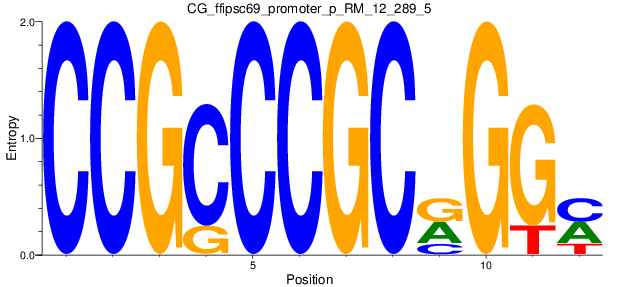 |
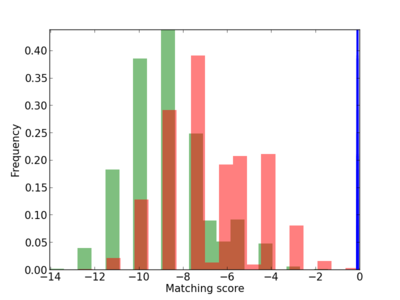 |
3 |
0.4 |
3.21E-04 |
MAVS |
positive regulation of IP-10 production [BP] |
| CG_ffipsc69_promoter_p_RM_12_12_7 |
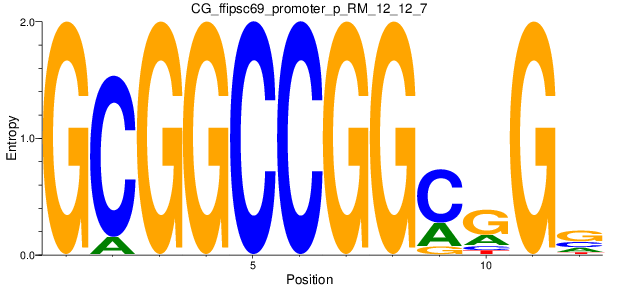 |
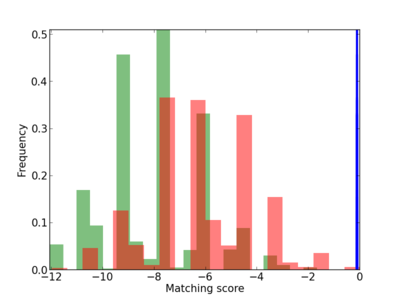 |
4 |
0.8 |
3.07E-04 |
GARNL3,
MAP4K4 |
small GTPase regulator activity [MF] |
| CG_ffipsc69_promoter_p_RM_16_1_9 |
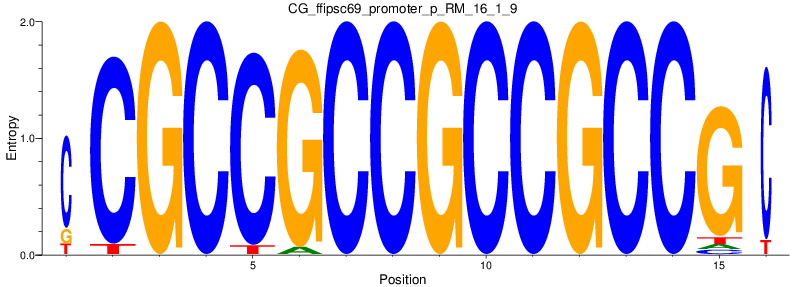 |
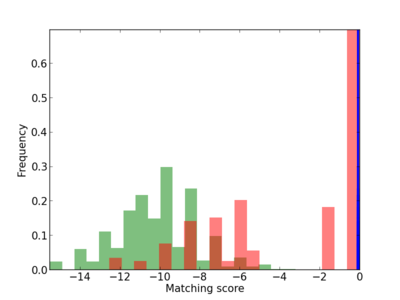 |
11 |
1.0 |
4.59E-04 |
GNAS,
NELFB,
ARID1A,
EGR1,
EPO,
BMPR2,
KCNIP3,
PKNOX1,
ACVR2A |
positive regulation of osteoblast differentiation [BP],
activin-activated receptor activity [MF],
response to hyperoxia [BP],
transforming growth factor beta-activated receptor activity [MF],
transmembrane receptor protein serine/threonine kinase activity [MF],
transcription from RNA polymerase II promoter [BP],
regulation of transcription from RNA polymerase II promoter [BP] |
| CG_ffipsc69_promoter_p_RM_12_16_11 |
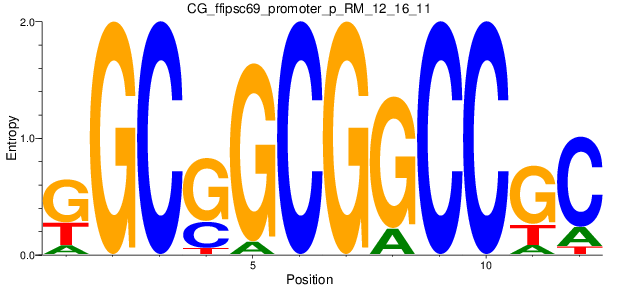 |
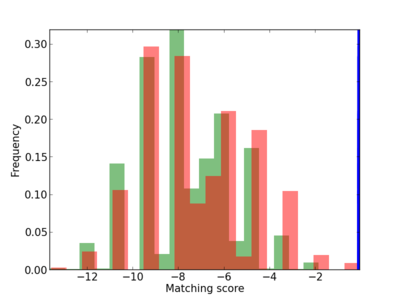 |
4 |
0.8 |
2.77E-04 |
TACC3,
TFAP2C |
dichotomous subdivision of terminal units involved in mammary gland duct morphogenesis [BP],
regulation of epidermis development [BP],
hair follicle development [BP],
cerebral cortex development [BP] |
| CG_ffipsc69_promoter_p_RM_12_20_9 |
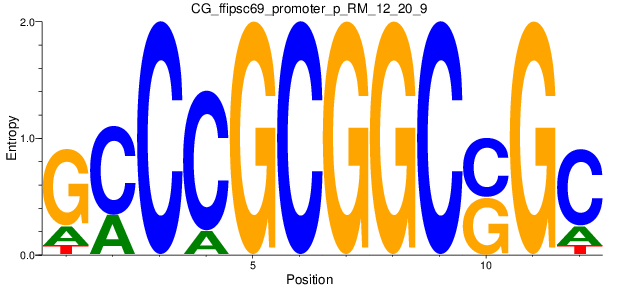 |
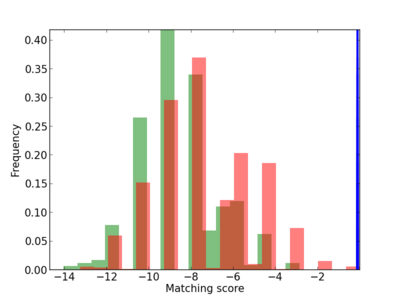 |
4 |
0.6 |
2.78E-04 |
PNMT,
LYN |
alpha2-beta1 integrin complex [CC],
phenylethanolamine N-methyltransferase activity [MF],
positive regulation of dendritic cell apoptotic process [BP],
negative regulation of mast cell proliferation [BP],
positive regulation of oligodendrocyte progenitor proliferation [BP] |
| CG_ffipsc69_promoter_p_RM_12_144_7 |
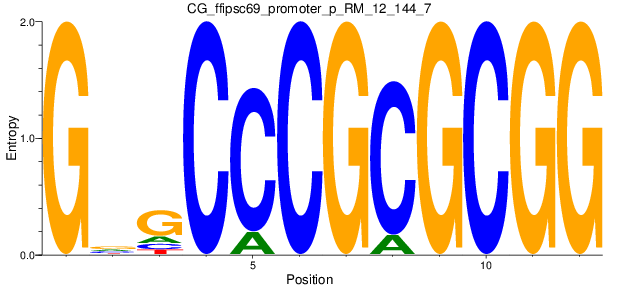 |
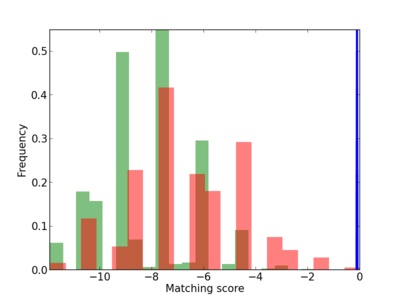 |
4 |
0.8 |
1.46E-04 |
NTMT1,
EGR1 |
N-terminal peptidyl-alanine trimethylation [BP],
N-terminal peptidyl-serine trimethylation [BP],
N-terminal peptidyl-serine dimethylation [BP],
positive regulation of glomerular metanephric mesangial cell proliferation [BP],
N-terminal peptidyl-proline dimethylation [BP] |
| CG_ffipsc69_promoter_p_RM_12_257_5 |
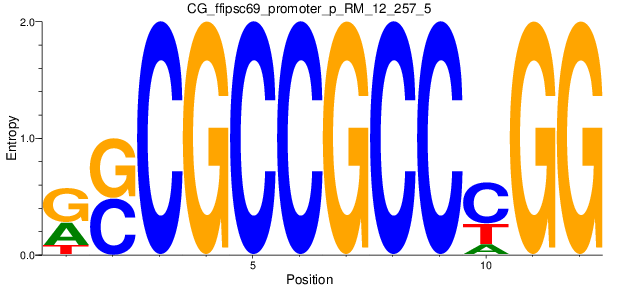 |
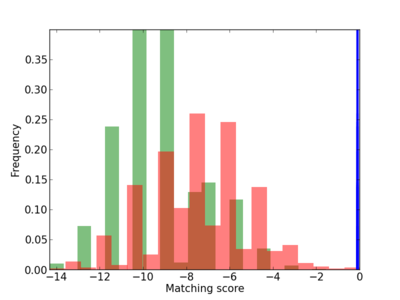 |
5 |
0.8 |
4.34E-04 |
SYK |
lymph vessel development [BP] |
| CG_ffipsc69_promoter_p_RM_12_253_3 |
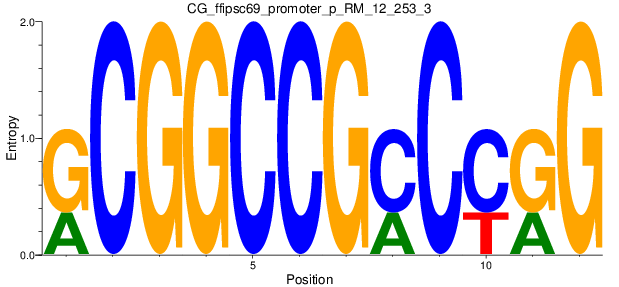 |
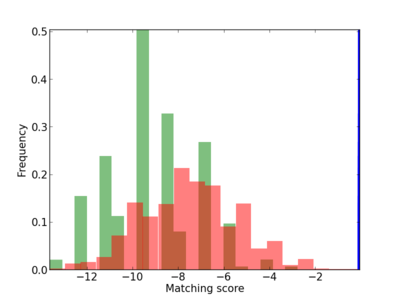 |
5 |
1.0 |
1.76E-04 |
DYNLL2,
HSPA14 |
synaptic target recognition [BP],
'de novo' cotranslational protein folding [BP] |
| CG_ffipsc69_promoter_p_RM_12_39_9 |
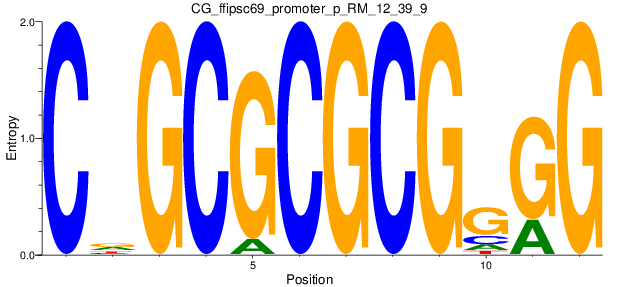 |
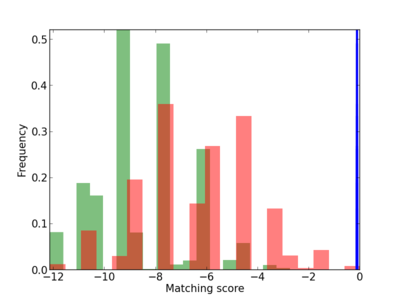 |
4 |
0.6 |
1.68E-04 |
BMP6,
SP5,
HOXC8,
RUNX2 |
bone morphogenesis [BP],
skeletal system morphogenesis [BP],
endochondral ossification [BP],
skeletal system development [BP] |
| CG_ffipsc69_promoter_p_RM_12_277_11 |
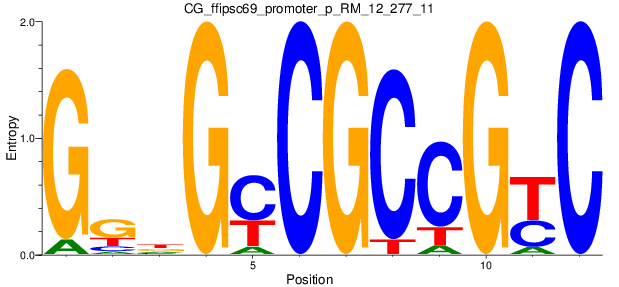 |
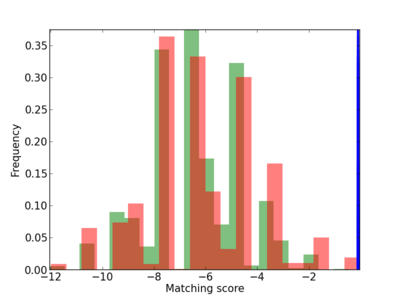 |
6 |
0.8 |
2.37E-04 |
GNPAT,
ADORA1 |
negative regulation of circadian sleep/wake cycle, non-REM sleep [BP],
negative regulation of mucus secretion [BP],
glycerone-phosphate O-acyltransferase activity [MF],
positive regulation of nucleoside transport [BP],
negative regulation of neurotrophin production [BP] |
| CG_ffipsc69_promoter_p_RM_16_8_1 |
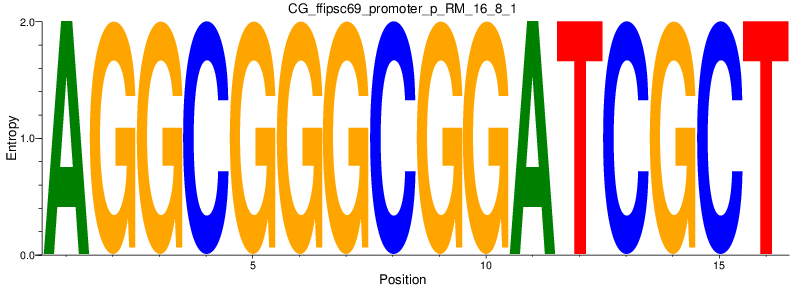 |
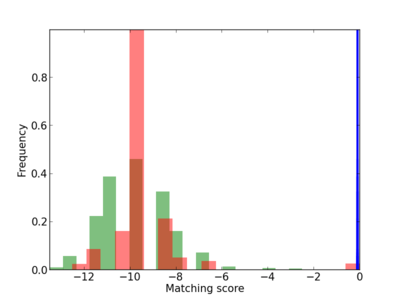 |
1 |
0.2 |
9.51E-05 |
ZDHHC17 |
magnesium ion transport [BP],
protein-cysteine S-palmitoleyltransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_129_8 |
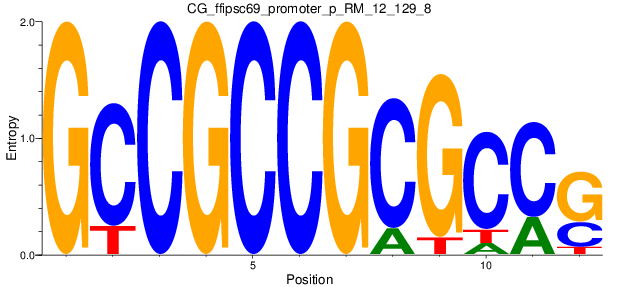 |
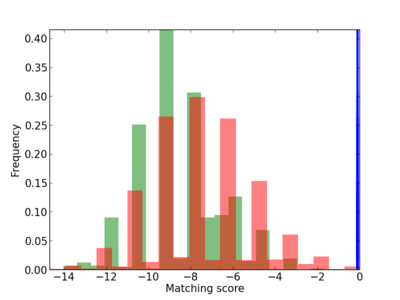 |
5 |
0.8 |
3.02E-04 |
STT3A |
co-translational protein modification [BP] |
| CG_ffipsc69_promoter_p_RM_12_14_9 |
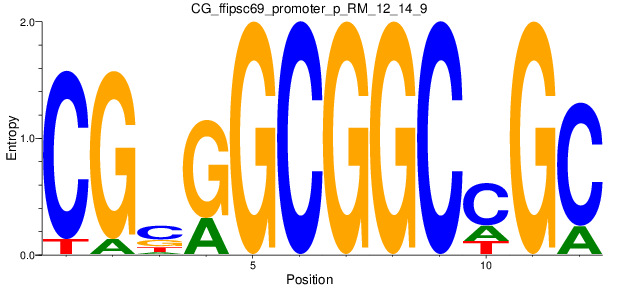 |
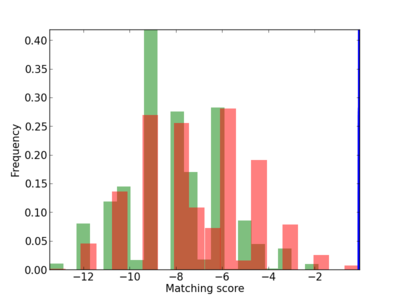 |
4 |
0.8 |
3.00E-04 |
CDC73,
BLOC1S3,
PI4K2A,
KIAA1967 |
RNA polymerase II core binding [MF],
BLOC-1 complex [CC] |
| CG_ffipsc69_promoter_p_RM_12_273_6 |
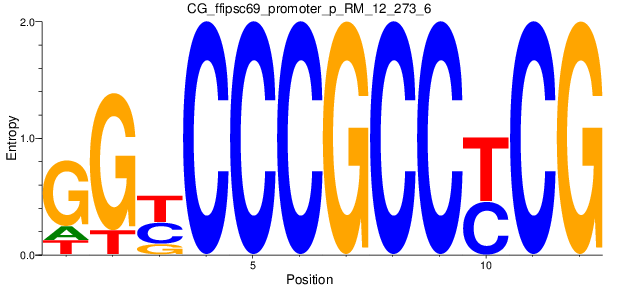 |
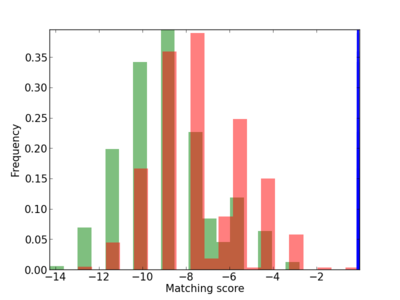 |
4 |
0.8 |
7.82E-05 |
FNTB |
protein farnesyltransferase complex [CC] |
| CG_ffipsc69_promoter_p_RM_12_142_4 |
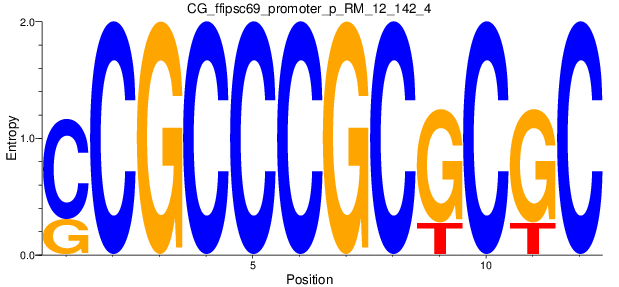 |
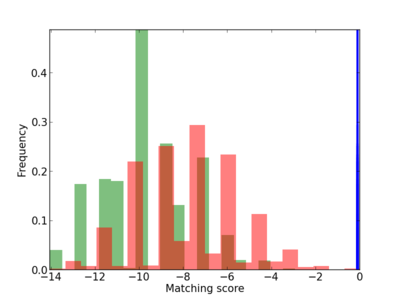 |
8 |
1.0 |
2.14E-04 |
AMOTL1,
DAO,
HEY2,
TEAD4,
LANCL2 |
D-alanine catabolic process [BP],
D-serine catabolic process [BP],
hippo signaling cascade [BP],
positive regulation of abscisic acid mediated signaling pathway [BP],
negative regulation of cardiac vascular smooth muscle cell differentiation [BP] |
| CG_ffipsc69_promoter_p_RM_12_41_13 |
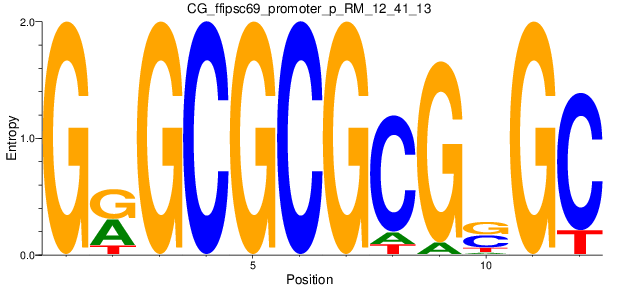 |
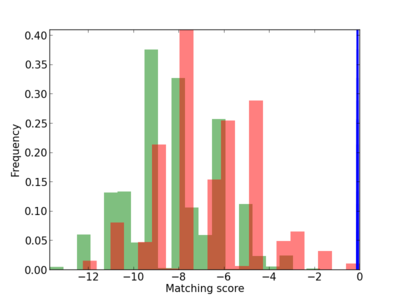 |
7 |
1.0 |
3.23E-04 |
PCNT,
MAPKAPK3 |
neural precursor cell proliferation [BP],
stress-activated MAPK cascade [BP] |
| CG_ffipsc69_promoter_p_RM_12_200_6 |
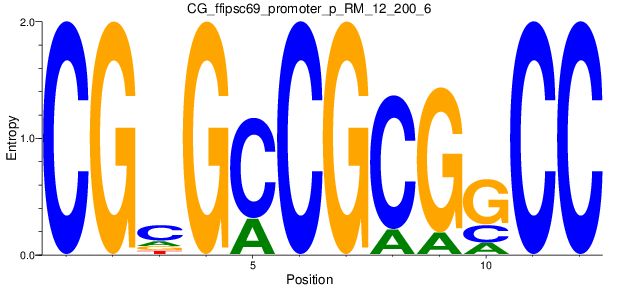 |
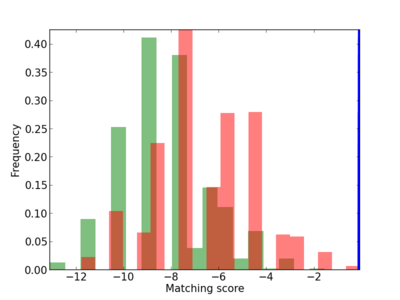 |
4 |
0.8 |
2.74E-04 |
USF2,
PTBP1,
SOX12,
MEIS1,
FOXJ2 |
sequence-specific DNA binding [MF],
transcription regulatory region sequence-specific DNA binding [MF],
RNA polymerase II distal enhancer sequence-specific DNA binding transcription factor activity [MF],
negative regulation of muscle cell differentiation [BP] |
| CG_ffipsc69_promoter_p_RM_16_3_2 |
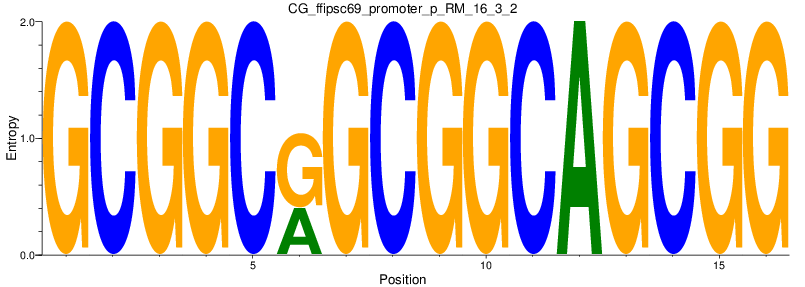 |
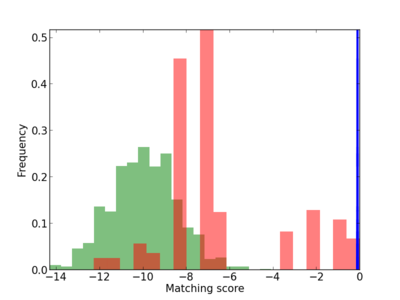 |
9 |
1.0 |
4.76E-04 |
AMD1 |
S-adenosylmethioninamine biosynthetic process [BP],
adenosylmethionine decarboxylase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_29_6 |
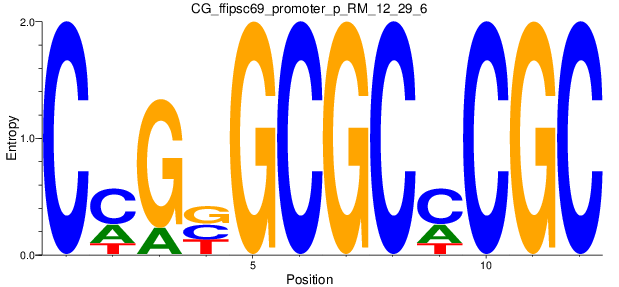 |
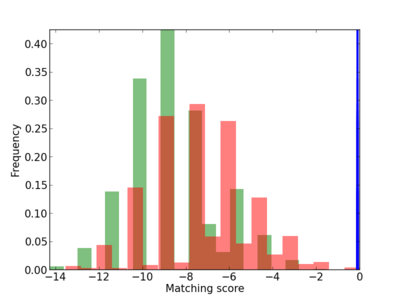 |
5 |
0.8 |
2.85E-04 |
HEY2 |
negative regulation of cardiac vascular smooth muscle cell differentiation [BP] |
| CG_ffipsc69_promoter_p_RM_12_117_6 |
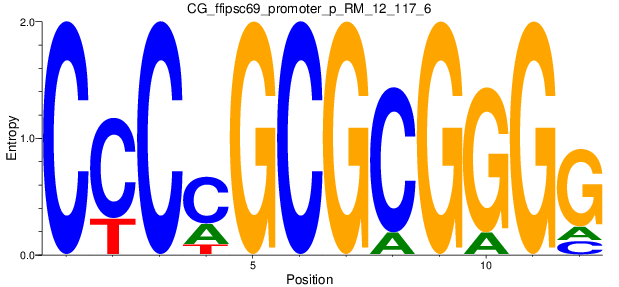 |
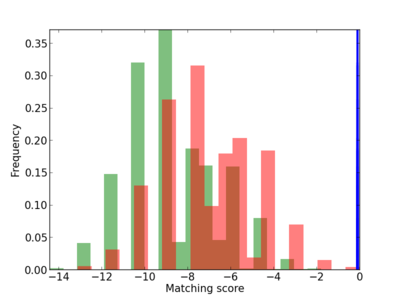 |
4 |
0.8 |
2.40E-04 |
ACSF3,
CHIA |
malonate catabolic process [BP],
cell wall chitin metabolic process [BP] |
| CG_ffipsc69_promoter_p_RM_12_76_8 |
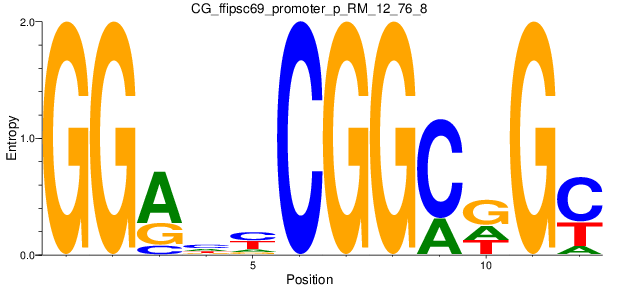 |
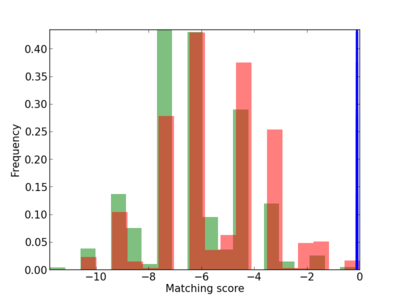 |
2 |
0.4 |
4.62E-04 |
HHEX,
LCN1,
GATA3,
GLYCTK |
negative regulation of fibroblast growth factor receptor signaling pathway involved in ureteric bud formation [BP],
sensory perception of taste [BP],
negative regulation of glial cell-derived neurotrophic factor receptor signaling pathway involved in ureteric bud formation [BP],
hepatic duct development [BP],
negative regulation of cell proliferation involved in mesonephros development [BP],
regulation of histone H3-K27 methylation [BP],
glycerate kinase activity [MF],
HMG box domain binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_10_5 |
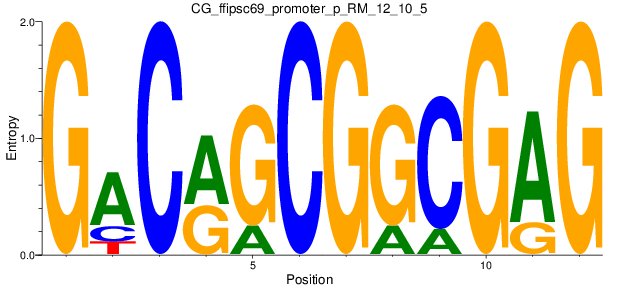 |
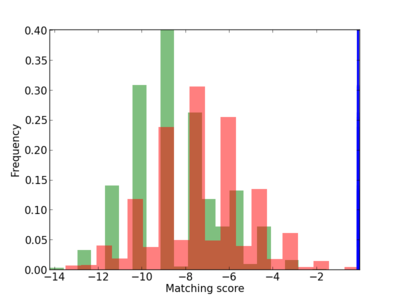 |
5 |
0.8 |
3.74E-04 |
MIB1,
ALMS1,
DYNC1H1,
RALGAPB |
centrosome [CC],
Ral protein signal transduction [BP] |
| CG_ffipsc69_promoter_p_RM_12_55_5 |
 |
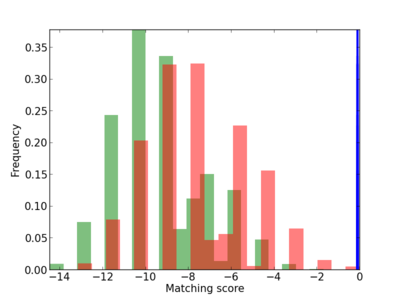 |
6 |
0.8 |
1.86E-04 |
KDM3A,
FOXL1,
OXT |
formaldehyde biosynthetic process [BP],
visceral mesoderm-endoderm interaction involved in midgut development [BP],
positive regulation of hindgut contraction [BP],
histone H3-K9 dimethylation [BP] |
| CG_ffipsc69_promoter_p_RM_12_199_5 |
 |
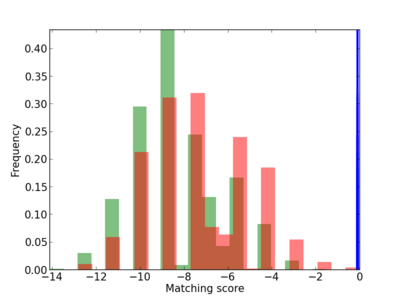 |
3 |
0.4 |
3.27E-04 |
MYC,
HOXD11,
SRR |
negative regulation of G2 phase of mitotic cell cycle [BP],
metanephros development [BP],
positive regulation of metanephric cap mesenchymal cell proliferation [BP],
D-serine ammonia-lyase activity [MF],
branching involved in ureteric bud morphogenesis [BP],
D-serine biosynthetic process [BP],
D-serine metabolic process [BP],
serine racemase activity [MF],
threonine racemase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_106_14 |
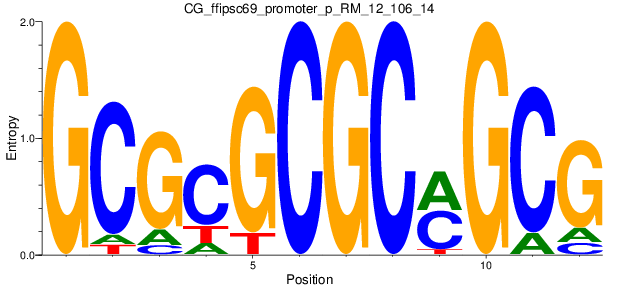 |
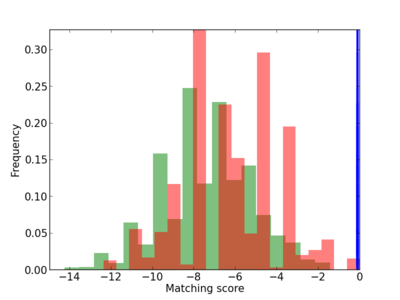 |
5 |
0.8 |
3.68E-04 |
SPHK1,
DOLPP1 |
sphingoid catabolic process [BP],
dolichyldiphosphatase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_271_4 |
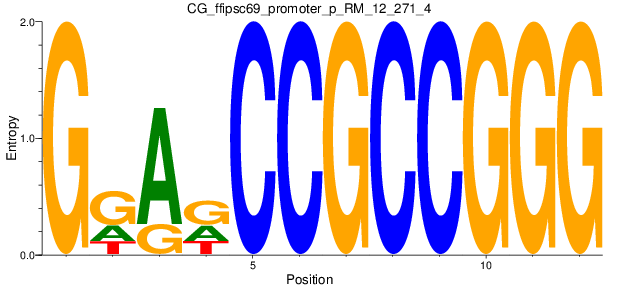 |
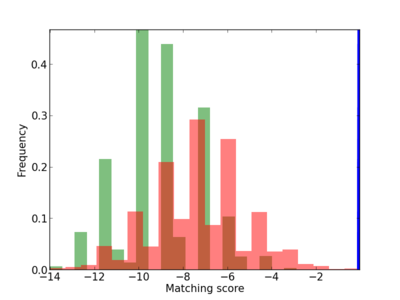 |
5 |
1.0 |
1.85E-04 |
IDUA,
CRHR1,
GATA4 |
lung lobe formation [BP],
corticotropin-releasing hormone receptor activity [MF],
L-iduronidase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_138_11 |
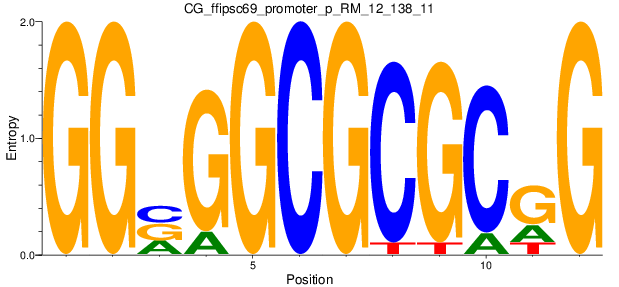 |
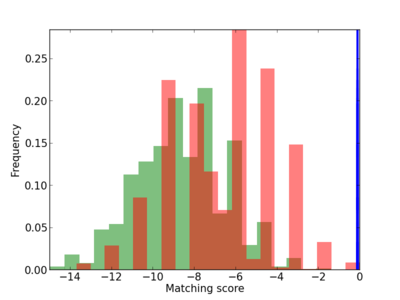 |
6 |
1.0 |
2.12E-04 |
SCN8A,
ATL1,
CHAT,
ADAM22,
SPTBN4 |
node of Ranvier [CC],
axon initial segment [CC],
axon [CC],
adult walking behavior [BP],
adult locomotory behavior [BP] |
| CG_ffipsc69_promoter_p_RM_12_148_11 |
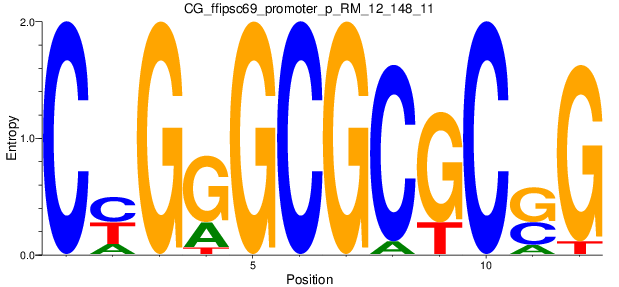 |
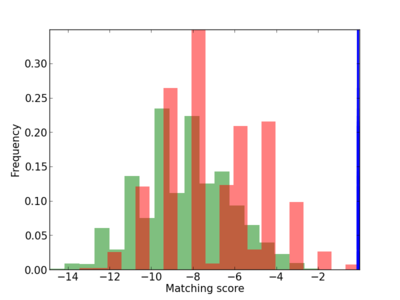 |
5 |
1.0 |
1.64E-04 |
TBX1,
CSF1,
BMPR1A |
regulation of ossification [BP],
odontogenesis of dentin-containing tooth [BP],
odontogenesis [BP],
positive regulation of mesenchymal cell proliferation [BP],
ossification [BP],
regulation of organ morphogenesis [BP],
heart formation [BP] |
| CG_ffipsc69_promoter_p_RM_12_235_7 |
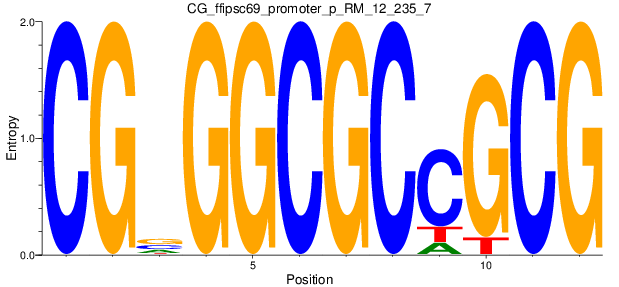 |
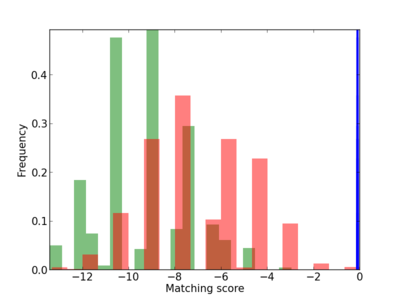 |
4 |
0.8 |
1.89E-04 |
SOCS6,
COMMD1 |
negative regulation of T cell activation [BP],
copper ion homeostasis [BP] |
| CG_ffipsc69_promoter_p_RM_14_6_2 |
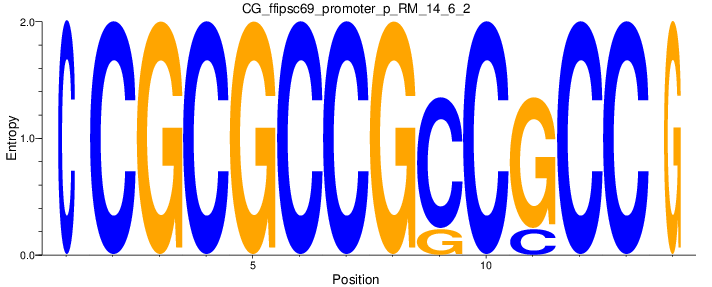 |
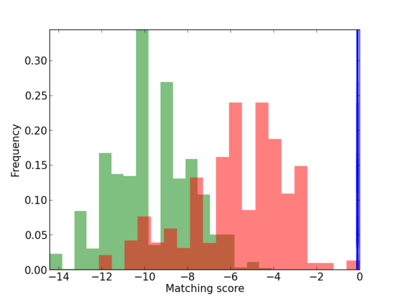 |
5 |
0.8 |
1.04E-04 |
HS3ST2,
FZD2 |
[heparan sulfate]-glucosamine 3-sulfotransferase 2 activity [MF],
muscular septum morphogenesis [BP] |
| CG_ffipsc69_promoter_p_RM_12_25_6 |
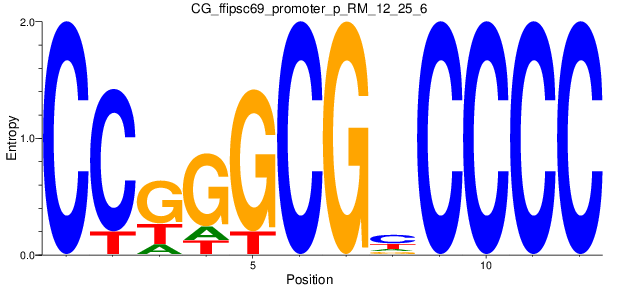 |
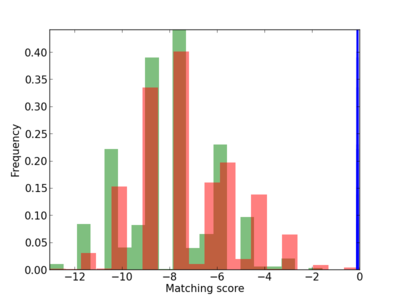 |
4 |
0.8 |
3.20E-04 |
OTX1,
BMPER,
GDF7 |
midbrain development [BP],
pathway-restricted SMAD protein phosphorylation [BP],
forebrain morphogenesis [BP],
regulation of pathway-restricted SMAD protein phosphorylation [BP] |
| CG_ffipsc69_promoter_p_RM_14_2_3 |
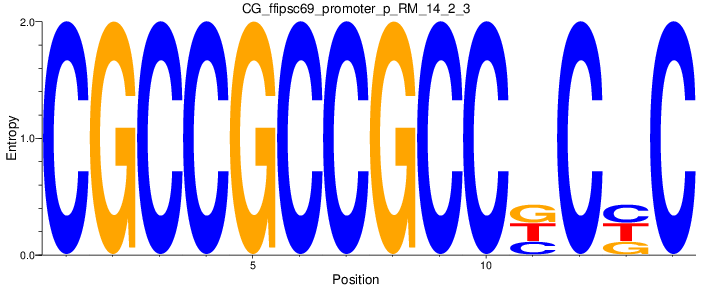 |
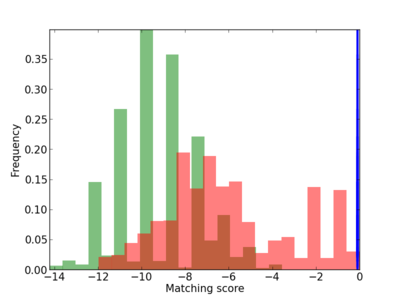 |
8 |
1.0 |
3.54E-04 |
MEPCE |
RNA methyltransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_115_6 |
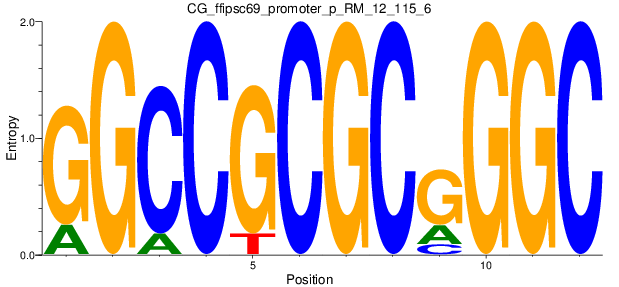 |
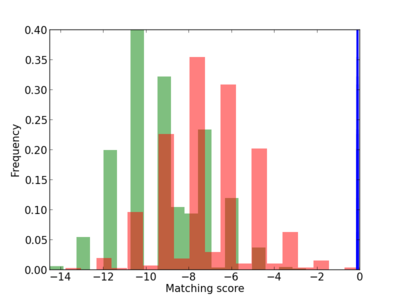 |
4 |
0.8 |
2.28E-04 |
PDZD3,
VIM,
DGKZ |
protein C-terminus binding [MF],
subapical complex [CC],
guanylate cyclase inhibitor activity [MF] |
| CG_ffipsc69_promoter_p_RM_16_7_1 |
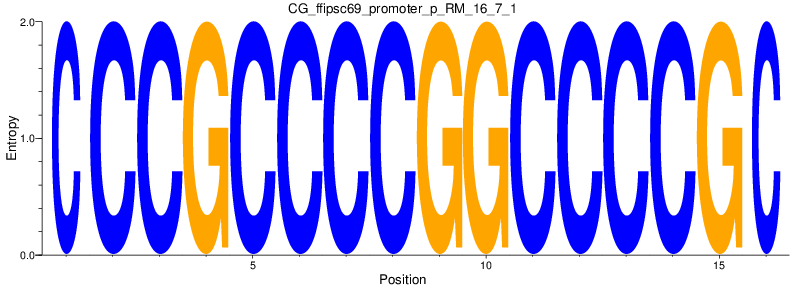 |
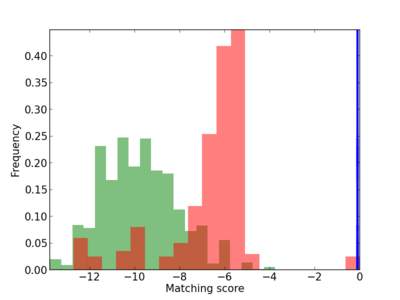 |
5 |
0.6 |
7.87E-05 |
RGCC,
SLC6A6 |
positive regulation of extracellular matrix constituent secretion [BP],
regulation of M/G1 transition of mitotic cell cycle [BP],
negative regulation of cell-cell adhesion mediated by cadherin [BP],
taurine:sodium symporter activity [MF],
negative regulation of exit from mitosis [BP],
negative regulation of fibroblast growth factor production [BP],
taurine binding [MF],
positive regulation of gene expression involved in extracellular matrix organization [BP],
taurine transport [BP] |
| CG_ffipsc69_promoter_p_RM_12_35_6 |
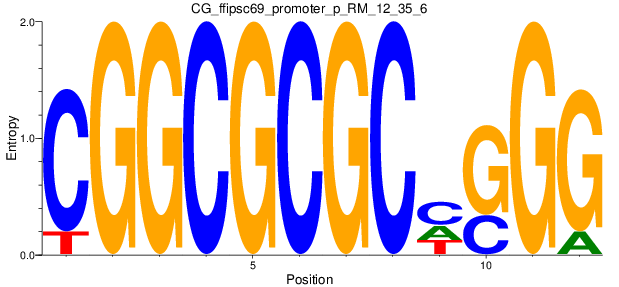 |
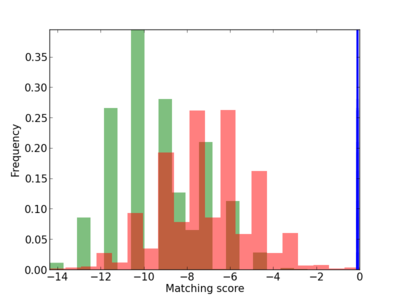 |
6 |
1.0 |
2.54E-04 |
PROX1 |
positive regulation of neural precursor cell proliferation [BP] |
| CG_ffipsc69_promoter_p_RM_12_28_5 |
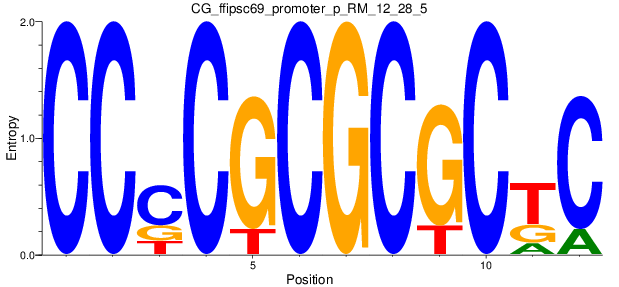 |
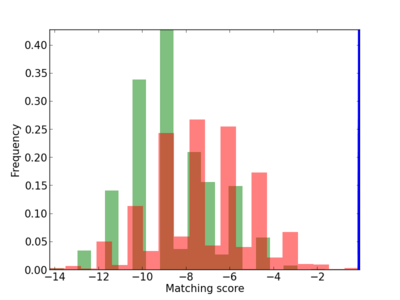 |
4 |
0.8 |
3.41E-04 |
AGK,
VEGFA,
NTRK1,
HSPB1,
SMAD4 |
positive regulation of blood vessel endothelial cell migration [BP],
neurotrophin p75 receptor binding [MF],
transforming growth factor beta receptor, common-partner cytoplasmic mediator activity [MF],
endothelial cell chemotaxis [BP],
basophil chemotaxis [BP],
positive regulation of endothelial cell chemotaxis by VEGF-activated vascular endothelial growth factor receptor signaling pathway [BP],
positive regulation of angiogenesis [BP],
positive regulation of endothelial cell migration [BP],
vascular endothelial growth factor receptor signaling pathway [BP],
cellular response to vascular endothelial growth factor stimulus [BP],
cellular response to growth factor stimulus [BP],
positive regulation of endothelial cell chemotaxis [BP],
acylglycerol kinase activity [MF] |
| CG_ffipsc69_promoter_p_RM_14_28_4 |
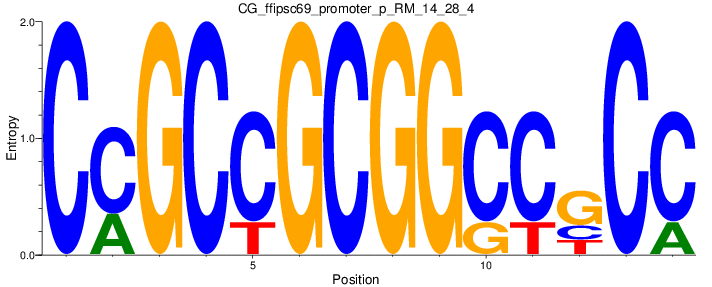 |
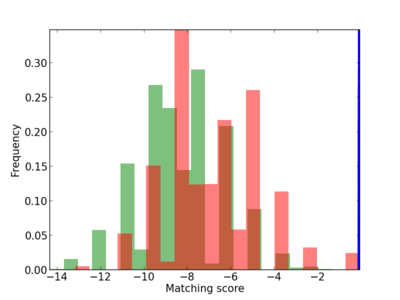 |
4 |
0.6 |
4.69E-04 |
ARID1A,
HTRA2,
CDK1 |
nucleosome mobilization [BP],
pentacyclic triterpenoid metabolic process [BP],
pronuclear fusion [BP] |
| CG_ffipsc69_promoter_p_RM_12_132_4 |
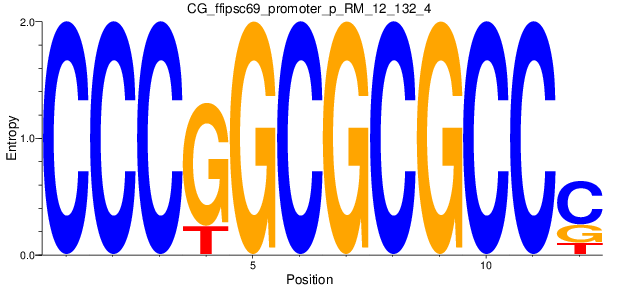 |
 |
5 |
1.0 |
2.43E-04 |
TCOF1,
HOXC10 |
skeletal system development [BP] |
| CG_ffipsc69_promoter_p_RM_12_275_7 |
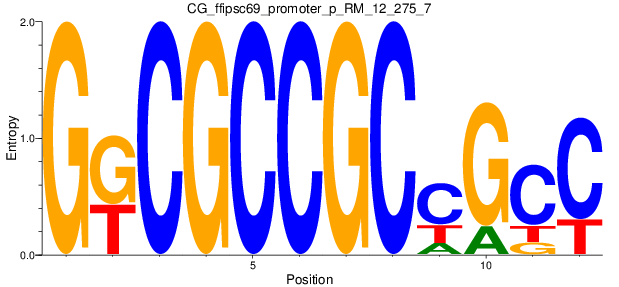 |
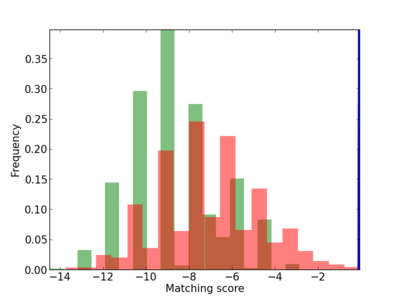 |
6 |
0.8 |
4.36E-04 |
GNAI2,
GNAZ,
LRP1 |
extrinsic to internal side of plasma membrane [CC],
lipoprotein transporter activity [MF],
G-protein beta/gamma-subunit complex binding [MF],
heterotrimeric G-protein complex [CC],
negative regulation of platelet-derived growth factor receptor-beta signaling pathway [BP],
protein complex binding [MF],
adenylate cyclase-inhibiting G-protein coupled receptor signaling pathway [BP] |
| CG_ffipsc69_promoter_p_RM_12_183_5 |
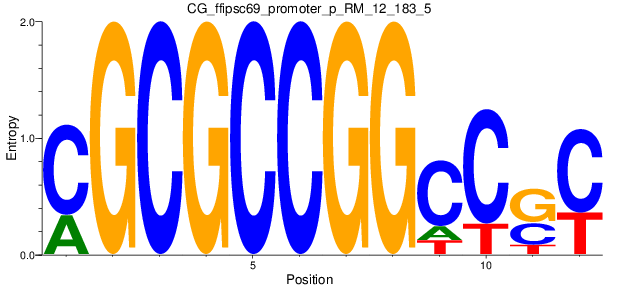 |
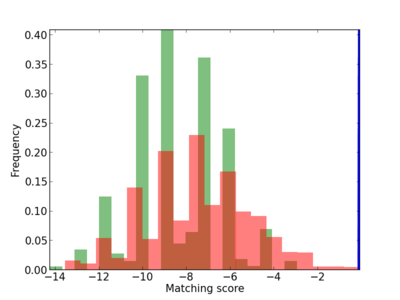 |
2 |
0.4 |
2.64E-04 |
MTHFD2,
TYMS |
tetrahydrofolate metabolic process [BP] |
| CG_ffipsc69_promoter_p_RM_16_5_1 |
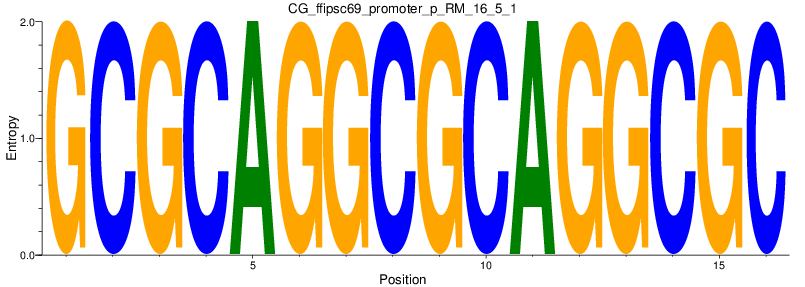 |
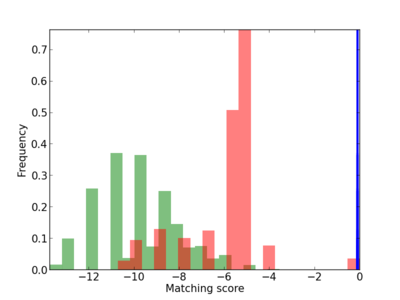 |
2 |
0.4 |
1.77E-04 |
DHTKD1 |
oxoglutarate dehydrogenase (succinyl-transferring) activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_60_3 |
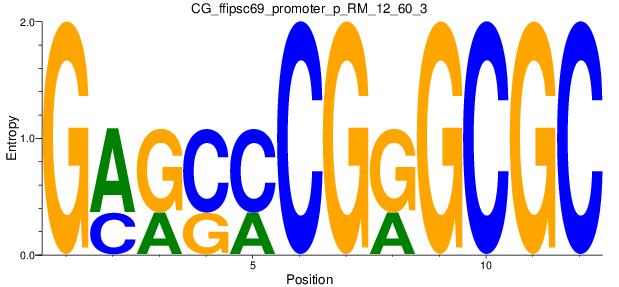 |
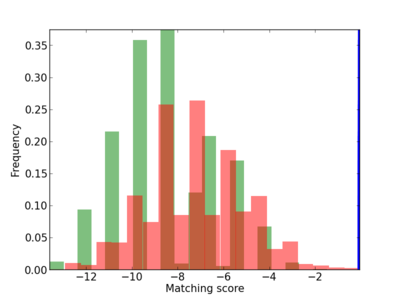 |
5 |
1.0 |
2.63E-04 |
GAL,
STUB1 |
regulation of glucocorticoid metabolic process [BP],
protein binding, bridging [MF] |
| CG_ffipsc69_promoter_p_RM_12_292_7 |
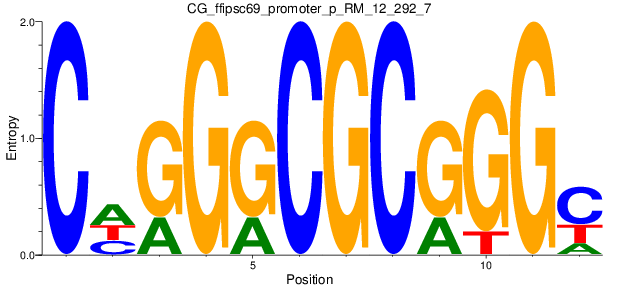 |
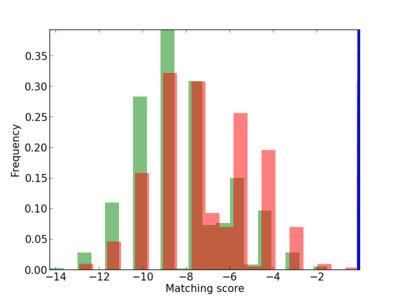 |
5 |
0.8 |
2.04E-04 |
TRMT61A |
tRNA (adenine-N1-)-methyltransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_14_18_3 |
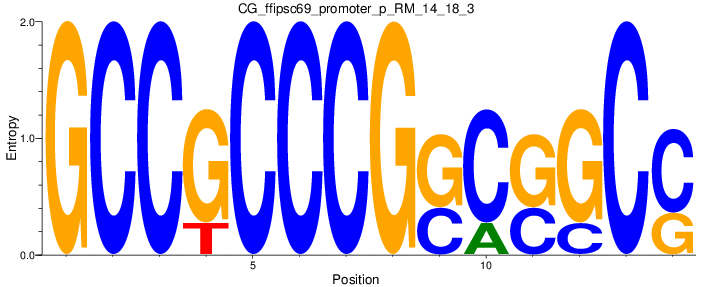 |
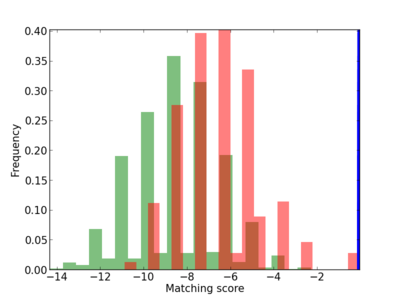 |
2 |
0.4 |
3.64E-04 |
ADCY1 |
response to lithium ion [BP] |
| CG_ffipsc69_promoter_p_RM_12_171_2 |
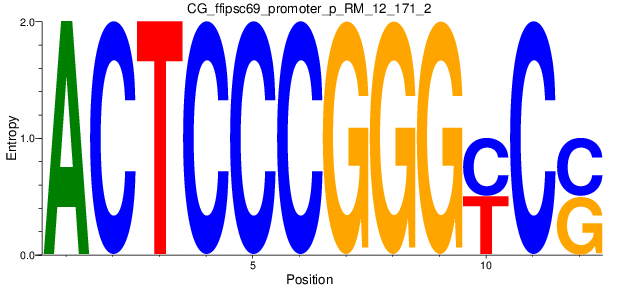 |
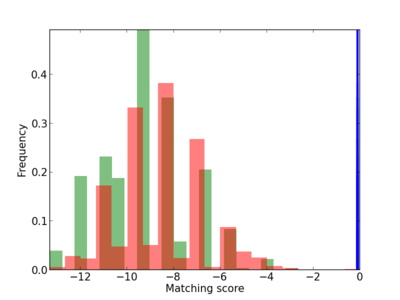 |
2 |
0.4 |
3.12E-04 |
PRKCG,
ARID1A |
nucleosome mobilization [BP],
positive regulation of mismatch repair [BP] |
| CG_ffipsc69_promoter_p_RM_12_233_8 |
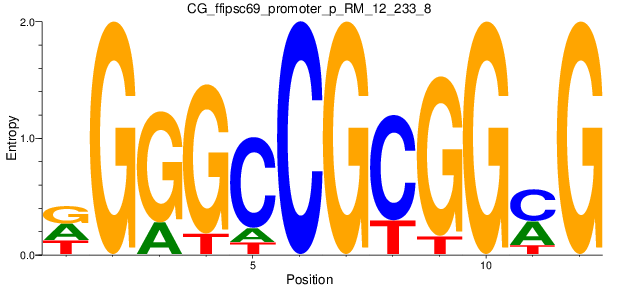 |
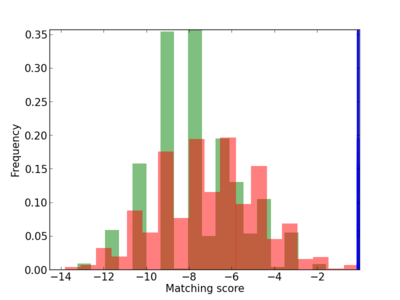 |
5 |
0.8 |
4.67E-04 |
BMP2,
RNASEH2A |
corticotropin hormone secreting cell differentiation [BP],
BMP signaling pathway involved in heart induction [BP],
positive regulation of Wnt receptor signaling pathway by BMP signaling pathway [BP],
mesenchymal cell proliferation involved in ureteric bud development [BP],
embryonic heart tube anterior/posterior pattern specification [BP],
ribonuclease H2 complex [CC],
negative regulation of cardiac muscle cell differentiation [BP] |
| CG_ffipsc69_promoter_p_RM_12_37_5 |
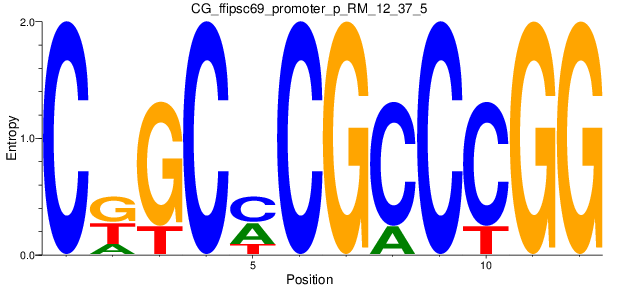 |
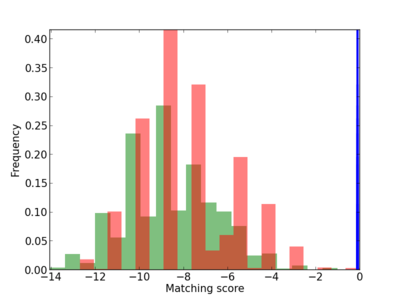 |
3 |
0.6 |
2.50E-04 |
AATF,
NR2E1,
SLC27A2,
BCAN |
negative regulation of superoxide anion generation [BP],
pristanate-CoA ligase activity [MF],
phytanate-CoA ligase activity [MF],
amygdala development [BP],
hippocampus development [BP] |
| CG_ffipsc69_promoter_p_RM_12_42_5 |
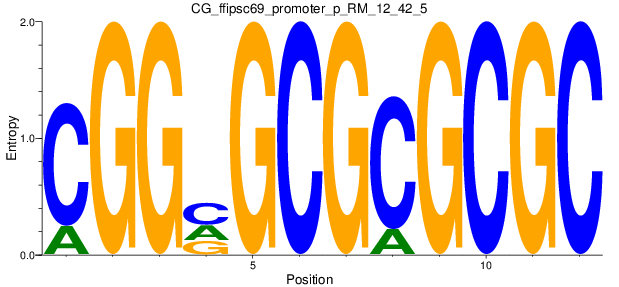 |
 |
5 |
0.8 |
4.78E-04 |
VAX2,
ZIC2 |
chromatin DNA binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_208_7 |
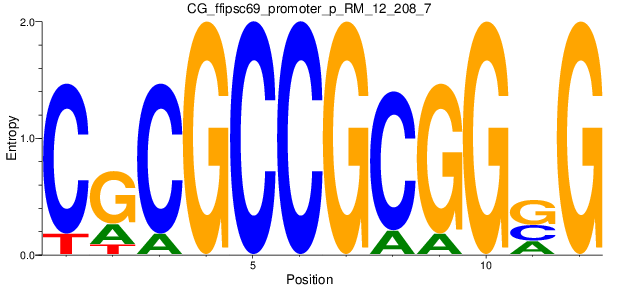 |
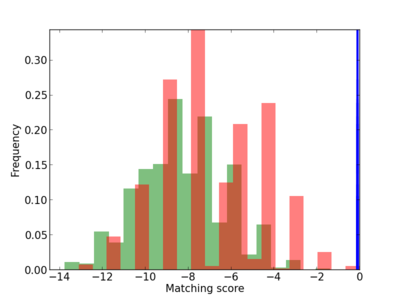 |
5 |
1.0 |
2.96E-04 |
ITSN1,
SIX3,
CYLD,
DAG1,
GIPR,
CDC73 |
Wnt receptor signaling pathway [BP],
nerve maturation [BP],
proline-rich region binding [MF],
cell surface receptor signaling pathway [BP],
protein import into nucleus [BP],
gastric inhibitory peptide receptor activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_172_3 |
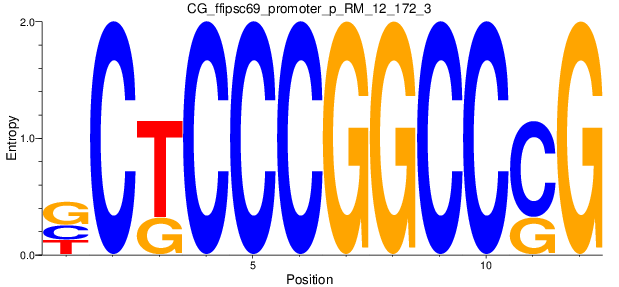 |
 |
3 |
0.6 |
1.19E-04 |
HEXB |
male courtship behavior [BP] |
| CG_ffipsc69_promoter_p_RM_12_81_13 |
 |
 |
4 |
0.8 |
4.16E-04 |
NFKBIB |
toll-like receptor signaling pathway [BP] |
| CG_ffipsc69_promoter_p_RM_18_2_2 |
 |
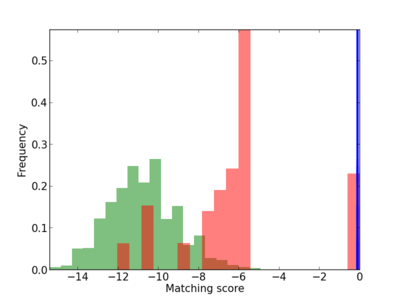 |
3 |
0.4 |
3.78E-04 |
ACER3 |
phytoceramidase activity [MF],
phytosphingosine biosynthetic process [BP] |
| CG_ffipsc69_promoter_p_RM_14_9_2 |
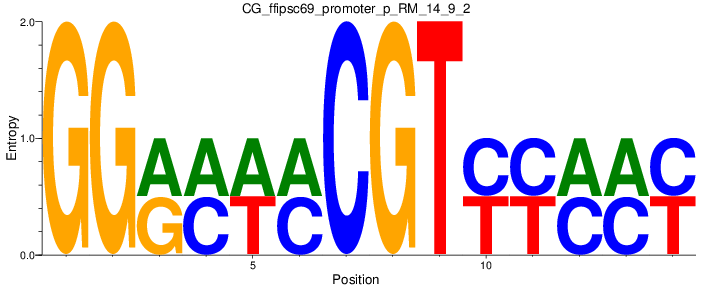 |
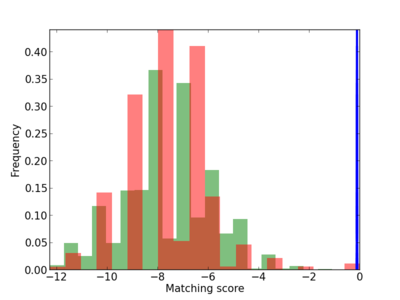 |
1 |
0.2 |
1.06E-04 |
TCOF1,
RBL2,
ZNF714,
ZNF85,
ZNF431,
ZNF430,
ZNF345,
ZNF493,
ZNF774,
BNIPL |
transcription, DNA-dependent [BP],
DNA binding [MF],
zinc ion binding [MF],
regulation of transcription, DNA-dependent [BP],
nucleus [CC] |
| CG_ffipsc69_promoter_p_RM_12_109_6 |
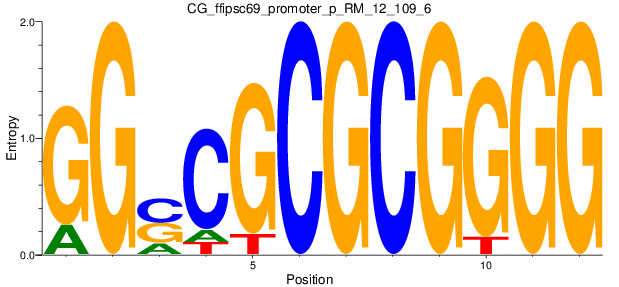 |
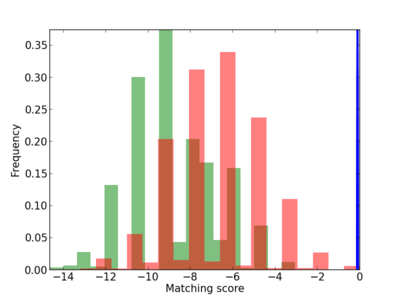 |
7 |
0.8 |
2.64E-04 |
DNASE1L2 |
endodeoxyribonuclease activity, producing 5'-phosphomonoesters [MF] |
| CG_ffipsc69_promoter_p_RM_12_120_6 |
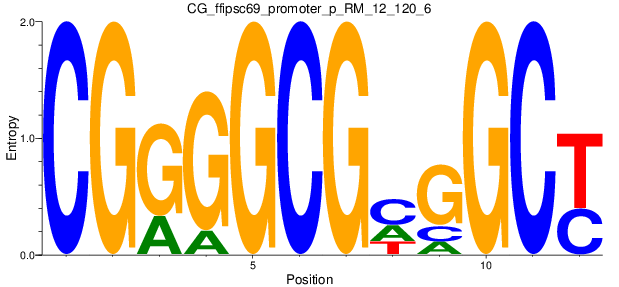 |
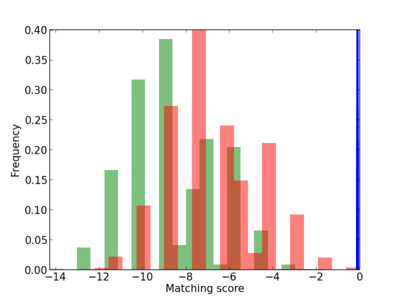 |
7 |
1.0 |
3.62E-04 |
NIT1,
F2RL1 |
positive regulation of eosinophil degranulation [BP],
positive regulation of renin secretion into blood stream [BP],
nitrilase activity [MF],
chemokine (C-C motif) ligand 2 secretion [BP],
positive regulation of neutrophil mediated killing of gram-negative bacterium [BP],
negative regulation of chemokine secretion [BP] |
| CG_ffipsc69_promoter_p_RM_12_64_5 |
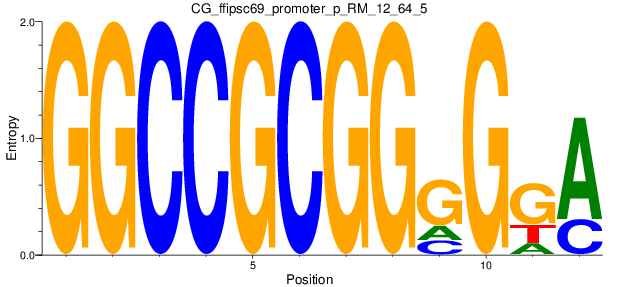 |
 |
4 |
0.8 |
3.09E-04 |
BMP2,
HOXD11,
NAT8L |
positive regulation of Wnt receptor signaling pathway by BMP signaling pathway [BP],
organ induction [BP],
aspartate N-acetyltransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_229_10 |
 |
 |
4 |
0.8 |
1.40E-04 |
DYNLL1,
RPS28,
HMGA1,
SLC12A5,
AATF,
FADD,
HTATIP2,
CRHR1,
IGF1R |
inactivation of MAPKK activity [BP],
G-protein alpha-subunit binding [MF],
viral reproduction [BP],
negative regulation of superoxide anion generation [BP],
thermosensory behavior [BP],
virus-host interaction [BP],
corticotropin-releasing hormone receptor activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_214_7 |
 |
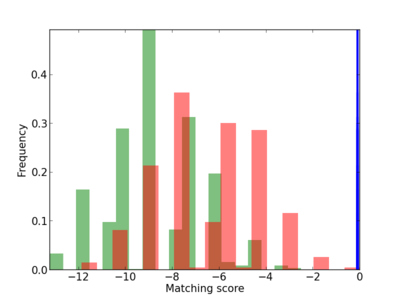 |
3 |
0.6 |
2.48E-04 |
RNASE2,
RNASE3 |
pancreatic ribonuclease activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_107_8 |
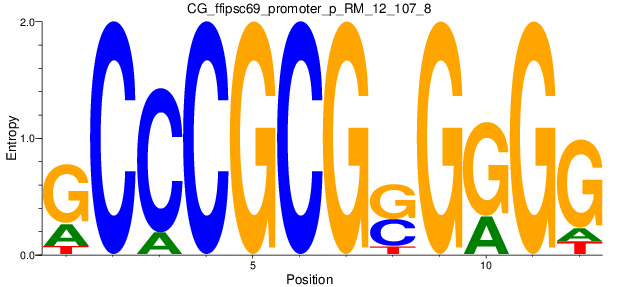 |
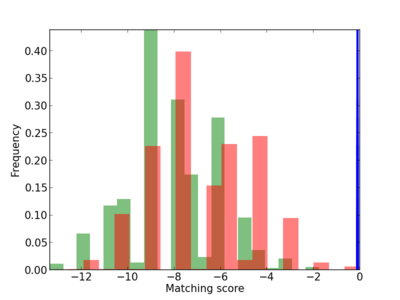 |
2 |
0.4 |
3.29E-04 |
BRDT |
histone displacement [BP],
positive regulation of transcription during meiosis [BP] |
| CG_ffipsc69_promoter_p_RM_12_113_6 |
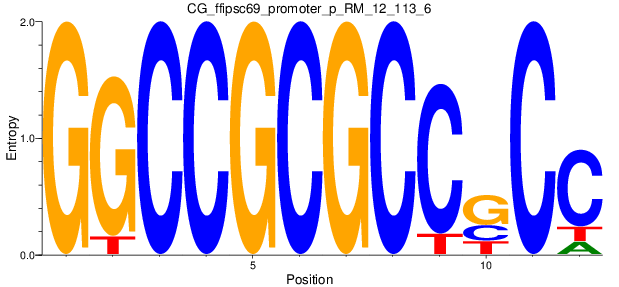 |
 |
4 |
0.8 |
3.59E-04 |
NR2E1,
MDK |
behavioral fear response [BP],
dentate gyrus development [BP] |
| CG_ffipsc69_promoter_p_RM_12_74_4 |
 |
 |
3 |
0.6 |
4.70E-04 |
GLYCTK,
FOXF2,
HHEX |
glycerate kinase activity [MF],
hepatic duct development [BP],
DNA binding, bending [MF] |
| CG_ffipsc69_promoter_p_RM_12_18_6 |
 |
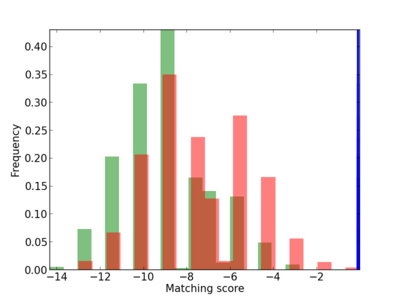 |
3 |
0.6 |
2.44E-04 |
DNM2,
LMTK2 |
transferrin transport [BP] |
| CG_ffipsc69_promoter_p_RM_18_4_1 |
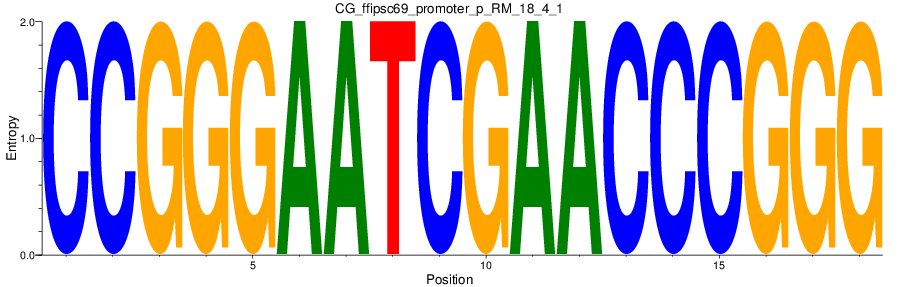 |
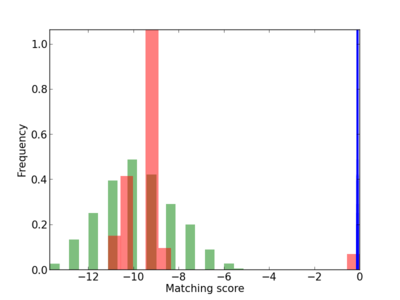 |
1 |
0.2 |
8.97E-04 |
PCYOX1 |
prenylcysteine metabolic process [BP],
chloride-transporting ATPase activity [MF],
prenylated protein catabolic process [BP],
prenylcysteine catabolic process [BP],
prenylcysteine oxidase activity [MF] |
| CG_ffipsc69_promoter_p_RM_14_36_2 |
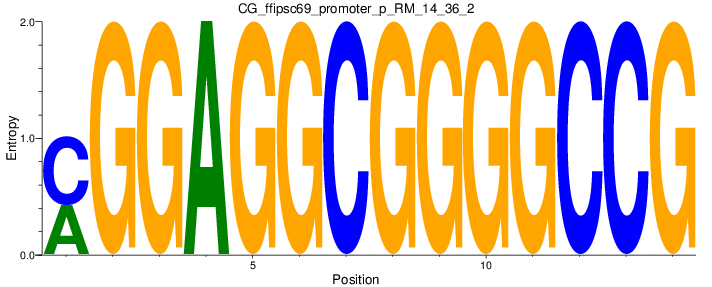 |
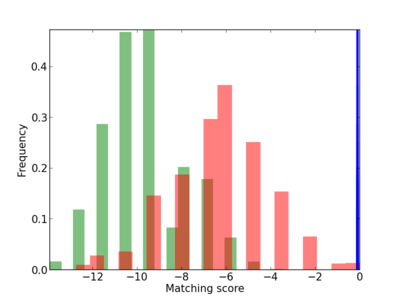 |
7 |
0.8 |
3.39E-04 |
YAP1,
TRPM4 |
positive regulation of canonical Wnt receptor signaling pathway [BP] |
| CG_ffipsc69_promoter_p_RM_14_20_3 |
 |
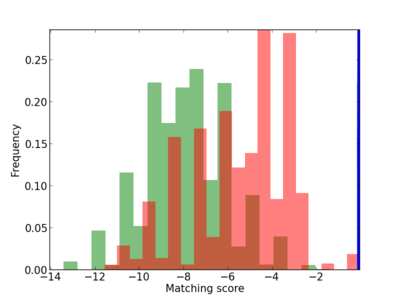 |
3 |
0.4 |
4.38E-04 |
GMPS,
GNPAT |
GMP synthase (glutamine-hydrolyzing) activity [MF],
GMP synthase activity [MF],
glycerone-phosphate O-acyltransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_195_7 |
 |
 |
2 |
0.4 |
3.37E-04 |
ISG20 |
exoribonuclease II activity [MF],
single-stranded DNA specific 3'-5' exodeoxyribonuclease activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_212_6 |
 |
 |
7 |
1.0 |
3.37E-04 |
PNP,
EGR1,
GALT |
NAD biosynthesis via nicotinamide riboside salvage pathway [BP],
purine-nucleoside phosphorylase activity [MF],
inosine catabolic process [BP],
nicotinamide riboside catabolic process [BP],
positive regulation of glomerular metanephric mesangial cell proliferation [BP],
UDP-glucose:hexose-1-phosphate uridylyltransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_173_4 |
 |
 |
3 |
0.6 |
2.87E-04 |
NGFR,
WNT10A |
hair follicle morphogenesis [BP],
regulation of odontogenesis of dentin-containing tooth [BP] |
| CG_ffipsc69_promoter_p_RM_12_285_6 |
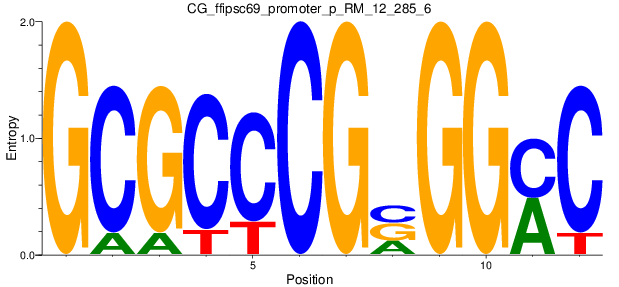 |
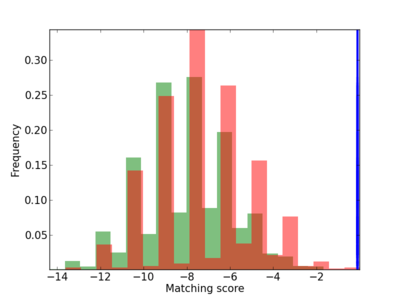 |
5 |
1.0 |
2.90E-04 |
USB1,
KANK1 |
U6 snRNA 3'-end processing [BP],
negative regulation of lamellipodium morphogenesis [BP] |
| CG_ffipsc69_promoter_p_RM_12_26_2 |
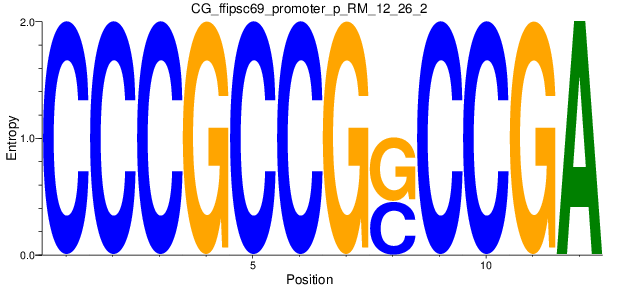 |
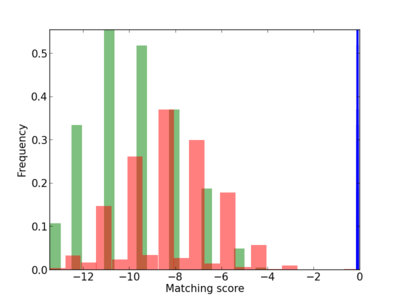 |
3 |
0.4 |
2.84E-05 |
PPP1CB,
LRSAM1,
PPOX |
oxygen-dependent protoporphyrinogen oxidase activity [MF],
myosin phosphatase activity [MF],
ubiquitin-dependent endocytosis [BP],
myosin-light-chain-phosphatase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_7_2 |
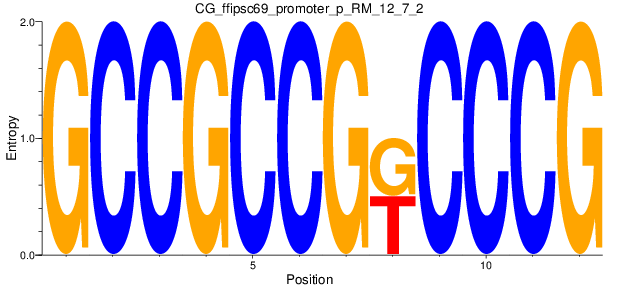 |
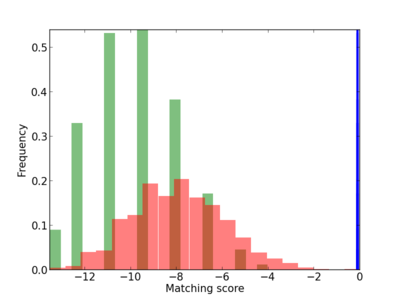 |
1 |
0.2 |
2.33E-04 |
STK11,
PTK2B |
3-phosphoinositide-dependent protein kinase binding [MF],
establishment of cell polarity [BP] |
| CG_ffipsc69_promoter_p_RM_12_158_4 |
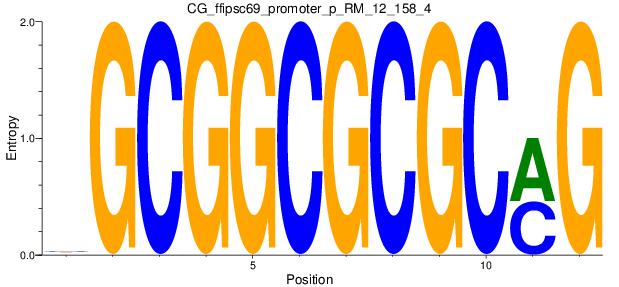 |
 |
4 |
0.8 |
3.73E-04 |
PROX1,
FOXG1 |
nose development [BP],
positive regulation of neural precursor cell proliferation [BP] |
| CG_ffipsc69_promoter_p_RM_12_207_6 |
 |
 |
4 |
0.6 |
2.38E-04 |
DOLPP1 |
dolichyldiphosphatase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_128_5 |
 |
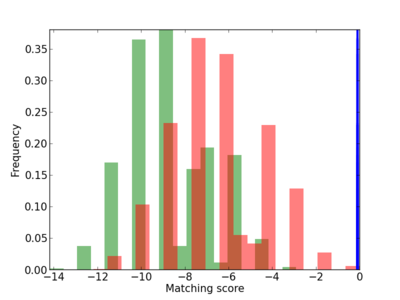 |
3 |
0.6 |
4.56E-04 |
UNCX,
ZIC2,
FOSL2,
ZNF135,
TEAD4,
AMOTL1 |
hippo signaling cascade [BP],
sequence-specific DNA binding transcription factor activity [MF] |
| CG_ffipsc69_promoter_p_RM_16_2_1 |
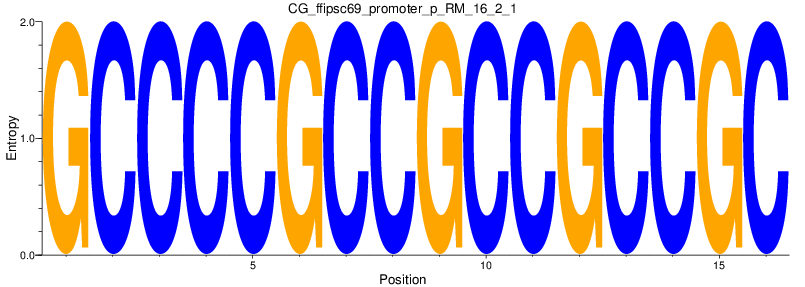 |
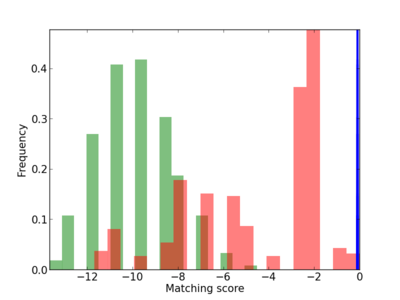 |
11 |
1.0 |
4.02E-04 |
USP43,
GNAS,
USP39 |
mu-type opioid receptor binding [MF],
post-embryonic body morphogenesis [BP],
ubiquitin thiolesterase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_131_5 |
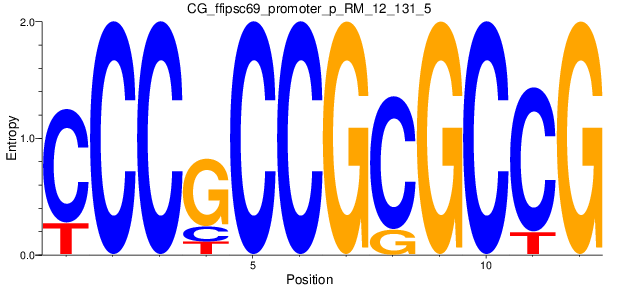 |
 |
5 |
1.0 |
4.14E-04 |
PRKCD,
WEE1 |
positive regulation of sphingomyelin catabolic process [BP],
negative regulation of filopodium assembly [BP],
positive regulation of phospholipid scramblase activity [BP],
non-membrane spanning protein tyrosine kinase activity [MF],
positive regulation of glucosylceramide catabolic process [BP] |
| CG_ffipsc69_promoter_p_RM_12_53_4 |
 |
 |
4 |
0.6 |
4.44E-04 |
ZNF76,
CTDSP1,
PKNOX2,
CBFB |
transcription from RNA polymerase II promoter [BP],
regulation of transcription from RNA polymerase II promoter [BP] |
| CG_ffipsc69_promoter_p_RM_14_5_3 |
 |
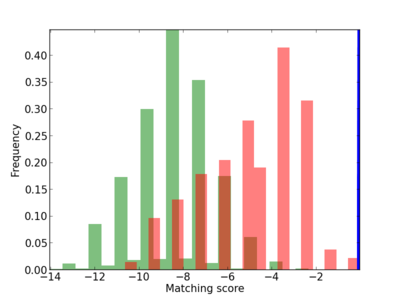 |
8 |
1.0 |
4.64E-04 |
GNAI3,
DERA,
GNAS,
PRMT6 |
extrinsic to internal side of plasma membrane [CC],
post-embryonic body morphogenesis [BP],
G-protein beta/gamma-subunit complex binding [MF],
histone methyltransferase activity (H3-R2 specific) [MF],
deoxyribose-phosphate aldolase activity [MF],
heterotrimeric G-protein complex [CC],
histone methyltransferase activity (H2A-R3 specific) [MF] |
| CG_ffipsc69_promoter_p_RM_12_108_10 |
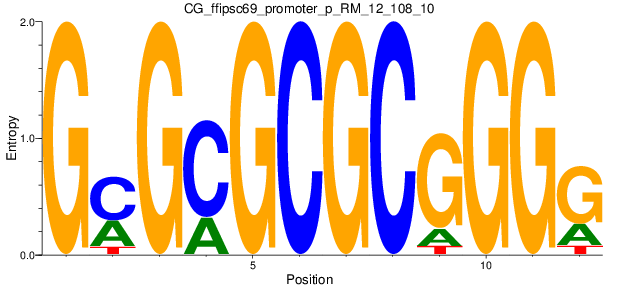 |
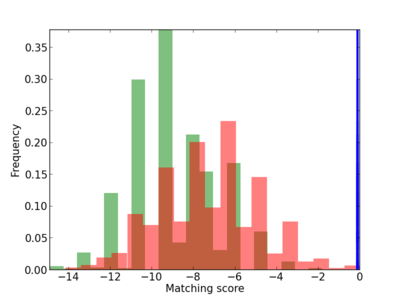 |
7 |
0.8 |
3.02E-04 |
RAB12,
ZFYVE27,
PLCB3,
PLCG2 |
phosphatidylinositol phospholipase C activity [MF],
recycling endosome membrane [CC] |
| CG_ffipsc69_promoter_p_RM_12_9_7 |
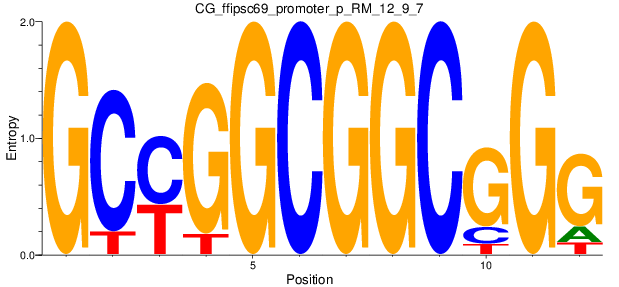 |
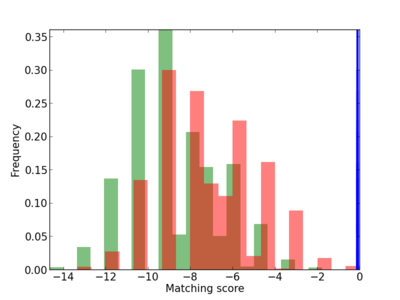 |
5 |
0.8 |
3.42E-04 |
CDC73,
PCM1 |
protein localization to centrosome [BP],
stem cell maintenance [BP],
positive regulation of Wnt receptor signaling pathway [BP] |
| CG_ffipsc69_promoter_p_RM_12_281_7 |
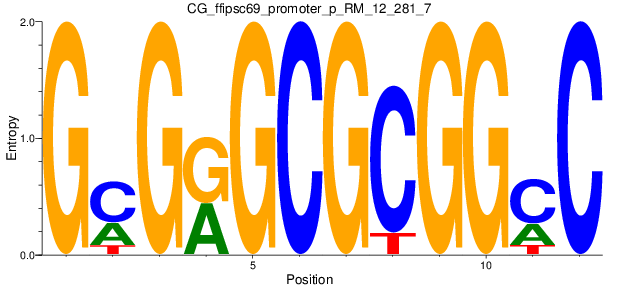 |
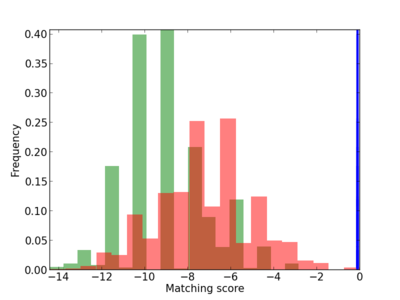 |
2 |
0.4 |
3.94E-04 |
KIAA1967,
NAGS,
MSH2,
MYO9B,
PDPK1,
CYP26A1,
ATP5C1 |
regulation of endothelial cell migration [BP],
ATP catabolic process [BP],
mitochondrial matrix [CC],
ADP binding [MF],
metabolic process [BP] |
| CG_ffipsc69_promoter_p_RM_14_7_1 |
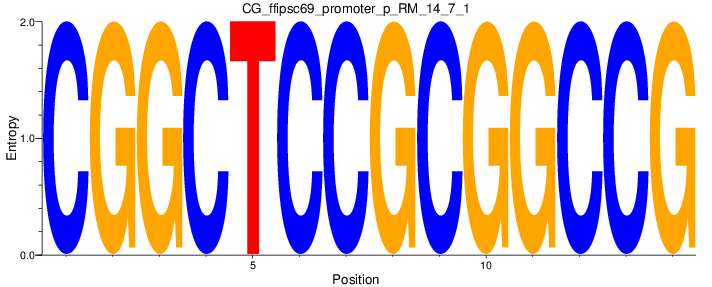 |
 |
3 |
0.4 |
3.95E-05 |
DNMT3B |
DNA (cytosine-5-)-methyltransferase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_224_9 |
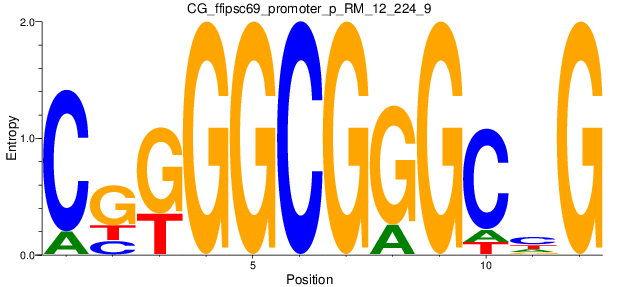 |
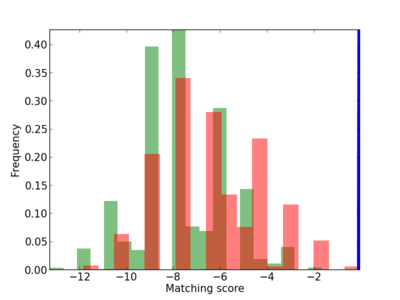 |
3 |
0.6 |
1.60E-04 |
YY1 |
response to prostaglandin F stimulus [BP] |
| CG_ffipsc69_promoter_p_RM_12_201_4 |
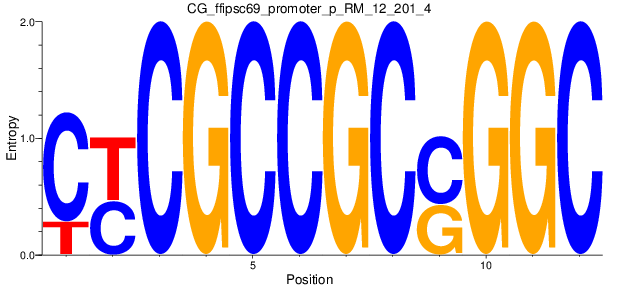 |
 |
6 |
1.0 |
1.29E-04 |
BCL2L11 |
BIM-BCL-2 complex [CC],
BIM-BCL-xl complex [CC] |
| CG_ffipsc69_promoter_p_RM_12_205_13 |
 |
 |
5 |
1.0 |
2.81E-04 |
EGR1,
LIAS,
FOXD2,
ITGA2,
CYP46A1,
NR2F1,
FZD2,
HOXD11,
AP2M1,
PKNOX1,
CREB3L1,
ZIC2,
SPTBN1,
TBX4,
AP2A2,
RNF4,
SIX6,
NRTN,
CTCF,
LHX2 |
axonogenesis [BP],
neural tube closure [BP],
neuron projection development [BP],
sequence-specific DNA binding transcription factor activity [MF],
sequence-specific DNA binding [MF],
axon guidance [BP],
morphogenesis of an epithelium [BP],
nervous system development [BP] |
| CG_ffipsc69_promoter_p_RM_12_101_4 |
 |
 |
5 |
0.8 |
2.37E-04 |
NUP85 |
lamellipodium assembly [BP] |
| CG_ffipsc69_promoter_p_RM_12_34_5 |
 |
 |
4 |
0.6 |
3.01E-04 |
LCK |
cellular zinc ion homeostasis [BP] |
| CG_ffipsc69_promoter_p_RM_14_30_2 |
 |
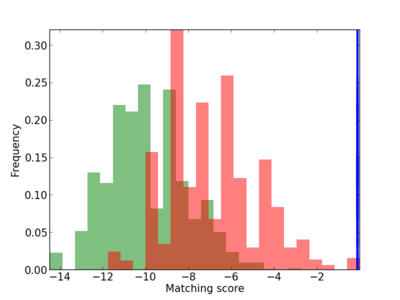 |
2 |
0.4 |
2.30E-04 |
MLL,
ADHFE1,
IQGAP1 |
neuron projection [CC],
positive regulation of cellular response to drug [BP],
hydroxyacid-oxoacid transhydrogenase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_196_5 |
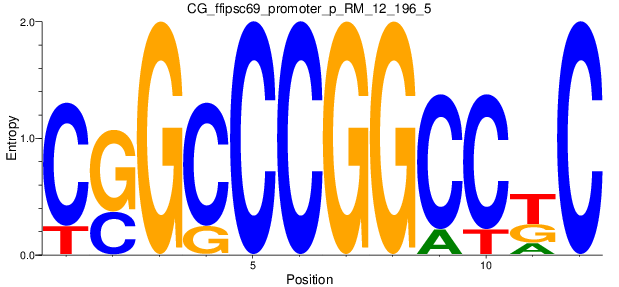 |
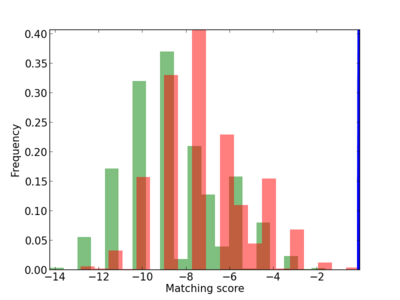 |
2 |
0.4 |
4.83E-04 |
RPS21,
FOXB1 |
mammillary body development [BP],
cell migration in diencephalon [BP],
inferior colliculus development [BP],
endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) [BP],
mammillothalamic axonal tract development [BP] |
| CG_ffipsc69_promoter_p_RM_12_291_5 |
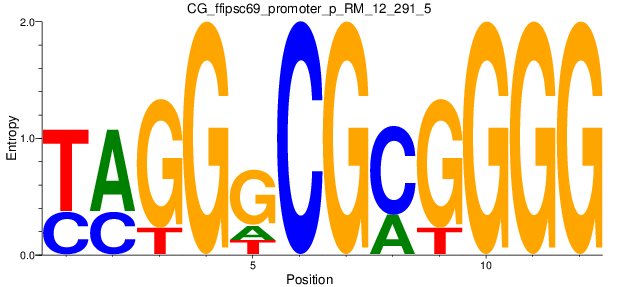 |
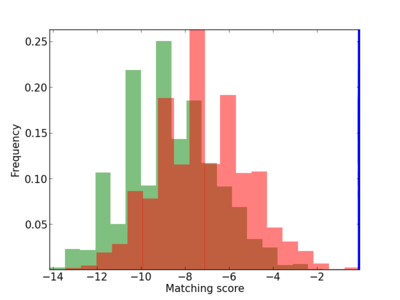 |
3 |
0.6 |
9.33E-05 |
MSH6,
PVRL2 |
sperm mitochondrion organization [BP],
virion attachment, binding of host cell surface coreceptor [BP],
single thymine insertion binding [MF],
oxidized purine DNA binding [MF],
susceptibility to T cell mediated cytotoxicity [BP],
MutSalpha complex [CC] |
| CG_ffipsc69_promoter_p_RM_12_87_13 |
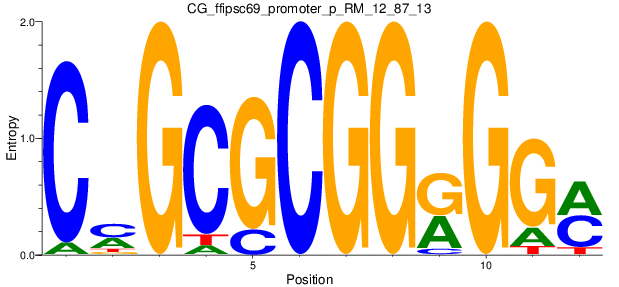 |
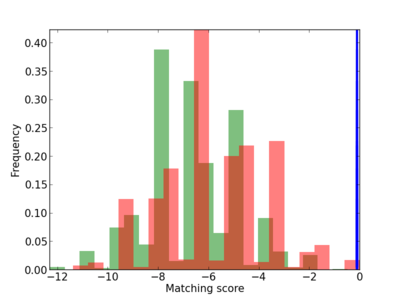 |
4 |
0.8 |
2.65E-04 |
NTMT1,
VEGFA |
N-terminal peptidyl-alanine trimethylation [BP],
N-terminal peptidyl-serine trimethylation [BP],
basophil chemotaxis [BP],
N-terminal peptidyl-serine dimethylation [BP],
N-terminal peptidyl-proline dimethylation [BP] |
| CG_ffipsc69_promoter_p_RM_18_3_1 |
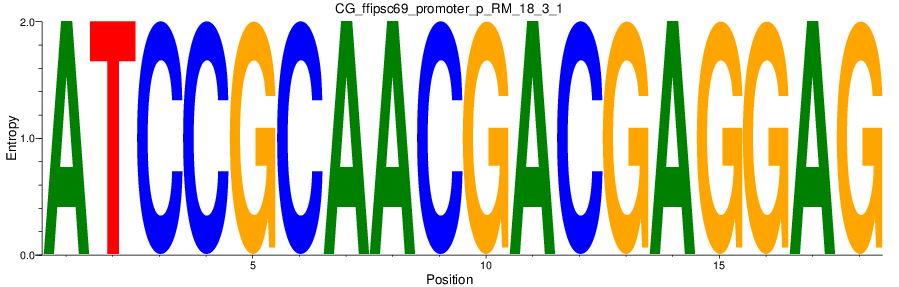 |
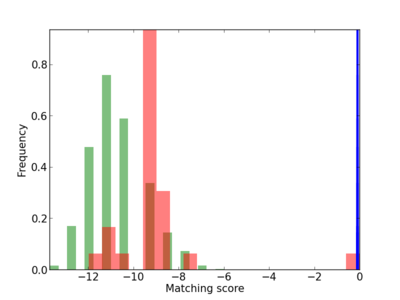 |
2 |
0.4 |
7.17E-04 |
HIST1H2AH,
HIST1H2AC,
HIST1H2BO |
nucleosome assembly [BP],
nucleosome [CC],
protein heterodimerization activity [MF] |
| CG_ffipsc69_promoter_p_RM_14_32_3 |
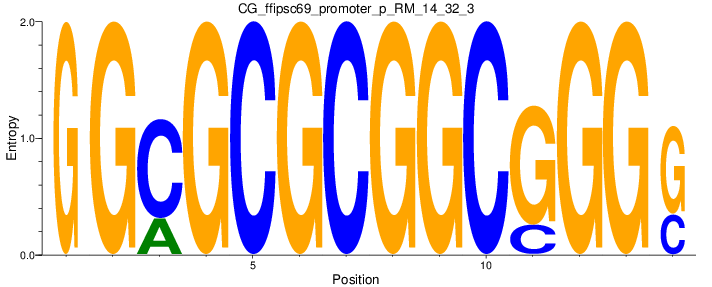 |
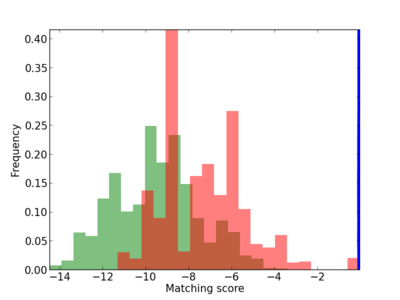 |
4 |
0.6 |
3.34E-04 |
MPP5 |
protein localization to myelin sheath abaxonal region [BP] |
| CG_ffipsc69_promoter_p_RM_12_165_7 |
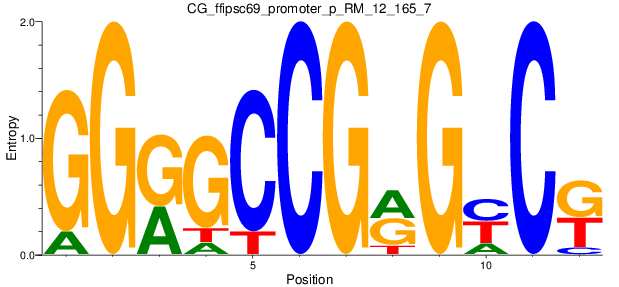 |
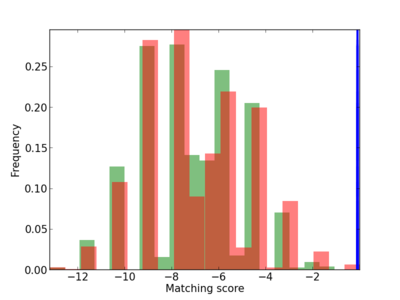 |
2 |
0.4 |
3.51E-04 |
PDX1 |
type B pancreatic cell differentiation [BP] |
| CG_ffipsc69_promoter_p_RM_12_230_4 |
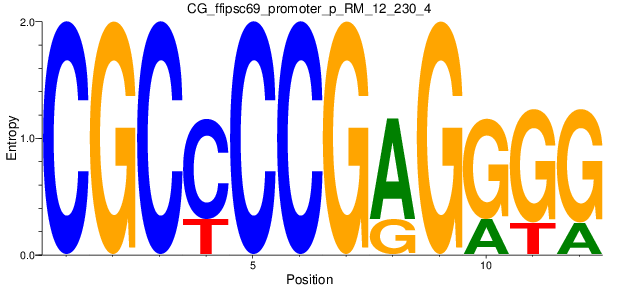 |
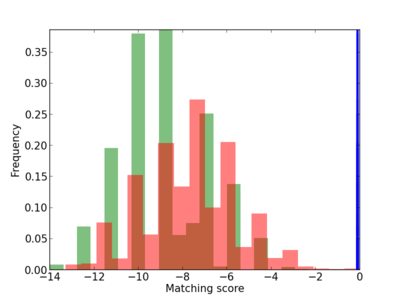 |
3 |
0.6 |
2.02E-04 |
CCNK,
CTF1 |
cyclin K-CDK13 complex [CC],
cyclin K-CDK12 complex [CC],
leukemia inhibitory factor receptor binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_272_10 |
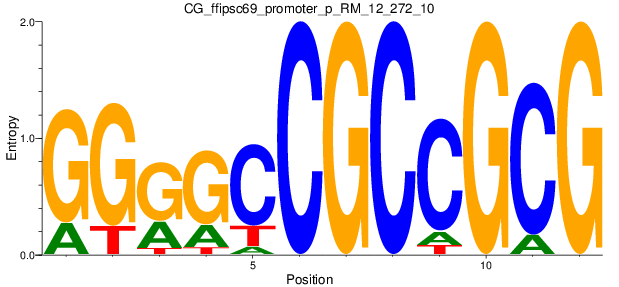 |
 |
5 |
1.0 |
3.27E-04 |
PGS1,
TLX2,
LMX1B,
HNRNPAB,
ZSCAN25,
SOX2,
NKX2-3,
GSX1 |
CDP-diacylglycerol-glycerol-3-phosphate 3-phosphatidyltransferase activity [MF],
detection of mechanical stimulus involved in equilibrioception [BP],
post-embryonic digestive tract morphogenesis [BP],
sequence-specific DNA binding transcription factor activity [MF],
sequence-specific DNA binding [MF],
adenohypophysis development [BP],
pituitary gland development [BP] |
| CG_ffipsc69_promoter_p_RM_12_151_8 |
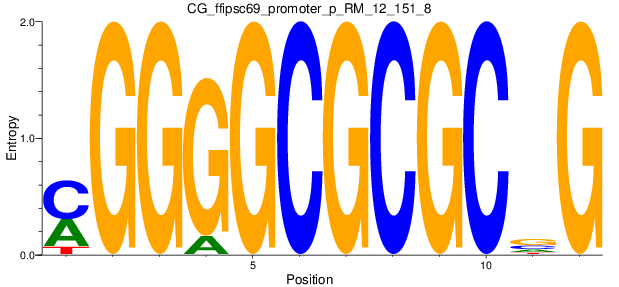 |
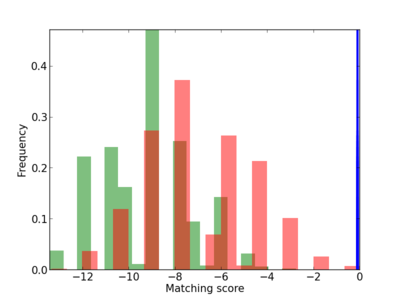 |
6 |
1.0 |
3.14E-04 |
DLX1,
HHEX,
HOXD8,
CIRH1A,
HOXD4,
SOX11,
MLLT10,
CBX2,
PTEN,
UTRN,
TEF,
FOXF2,
PAK6,
ZNF517,
TRAF3,
CDC42,
EDARADD |
forebrain morphogenesis [BP],
dystrophin-associated glycoprotein complex [CC],
odontogenesis of dentin-containing tooth [BP],
thioesterase binding [MF],
regulation of transcription, DNA-dependent [BP],
sequence-specific DNA binding transcription factor activity [MF],
transcription, DNA-dependent [BP],
sequence-specific DNA binding [MF],
establishment of planar polarity of embryonic epithelium [BP],
nucleic acid binding transcription factor activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_61_9 |
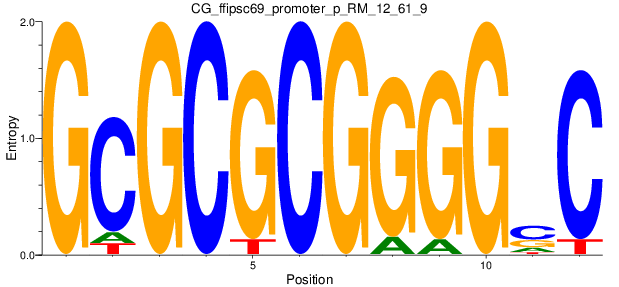 |
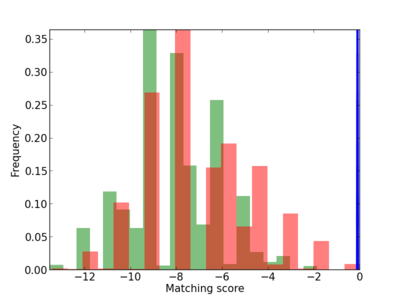 |
4 |
0.8 |
1.77E-04 |
SOX11,
NIPBL,
ZFPM2 |
embryonic digestive tract morphogenesis [BP],
RNA polymerase II transcription coactivator activity [MF],
outflow tract morphogenesis [BP] |
| CG_ffipsc69_promoter_p_RM_14_3_3 |
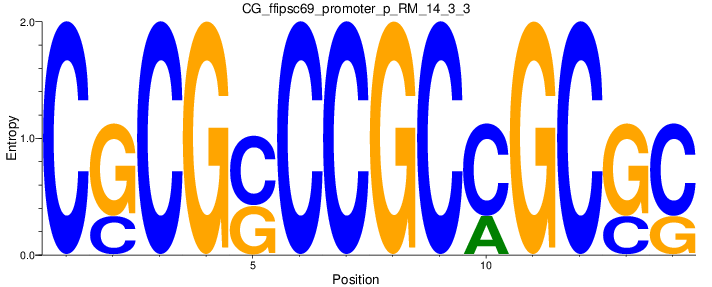 |
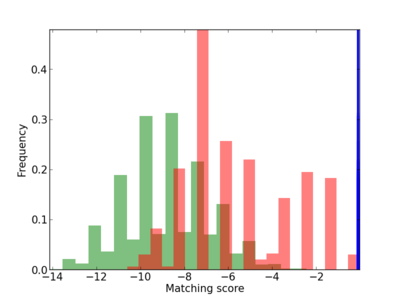 |
3 |
0.6 |
3.86E-04 |
PAX2,
DMRT3,
IL27RA,
HOXD11,
EN2 |
negative regulation of mesenchymal cell apoptotic process involved in metanephric nephron morphogenesis [BP],
negative regulation of apoptotic process involved in metanephric nephron tubule development [BP],
optic nerve structural organization [BP],
vestibulocochlear nerve formation [BP],
interleukin-27 receptor activity [MF],
positive regulation of optic nerve formation [BP],
optic chiasma development [BP],
negative regulation of mesenchymal cell apoptotic process involved in metanephros development [BP],
pronephric field specification [BP],
sequence-specific DNA binding [MF],
branching involved in ureteric bud morphogenesis [BP],
positive regulation of metanephric DCT cell differentiation [BP],
metanephros development [BP],
regulation of metanephros size [BP],
negative regulation of apoptotic process involved in metanephric collecting duct development [BP] |
| CG_ffipsc69_promoter_p_RM_12_99_6 |
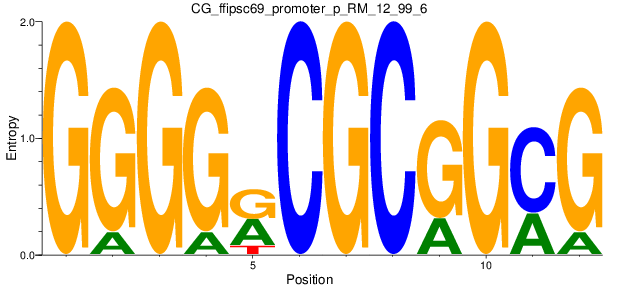 |
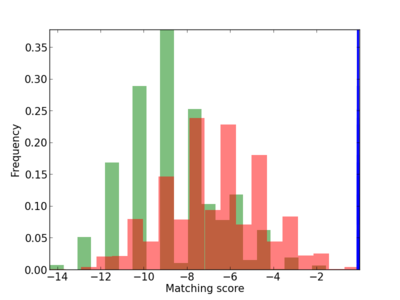 |
5 |
0.8 |
3.31E-04 |
POU3F3,
NOG,
RUNX2 |
negative regulation of cytokine activity [BP],
metanephric macula densa development [BP],
cellular response to BMP stimulus [BP],
fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation [BP],
regulation of fibroblast growth factor receptor signaling pathway [BP],
regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation [BP] |
| CG_ffipsc69_promoter_p_RM_12_280_8 |
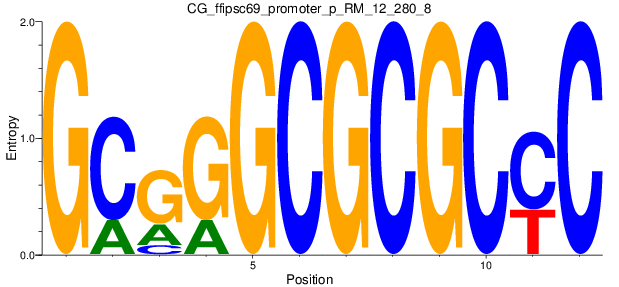 |
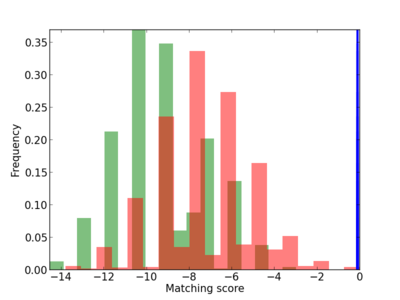 |
3 |
0.6 |
1.94E-04 |
ANGPTL4,
PRKCE,
CBX2,
BMP1,
BARX2 |
positive regulation of MAPK cascade [BP],
multicellular organismal development [BP],
development of primary sexual characteristics [BP],
cell differentiation [BP],
cartilage condensation [BP] |
| CG_ffipsc69_promoter_p_RM_12_88_2 |
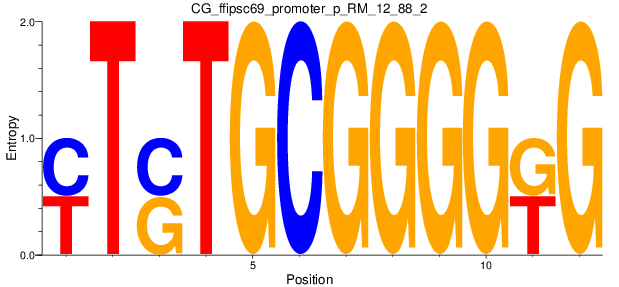 |
 |
2 |
0.4 |
1.76E-04 |
QKI,
P2RX2,
PDP2,
NAA60,
AGRN |
muscle cell differentiation [BP],
neuromuscular junction development [BP],
N-terminal peptidyl-methionine acetylation [BP],
magnesium-dependent protein serine/threonine phosphatase activity [MF],
detection of hypoxic conditions in blood by carotid body chemoreceptor signaling [BP],
positive regulation of synaptic growth at neuromuscular junction [BP] |
| CG_ffipsc69_promoter_p_RM_12_236_6 |
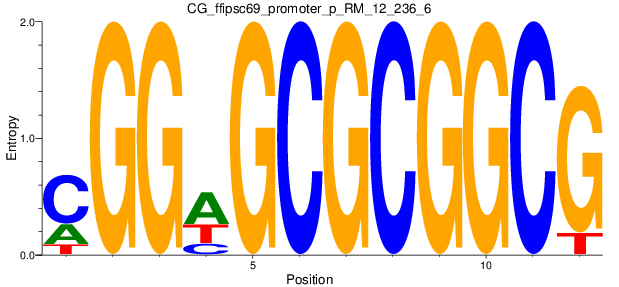 |
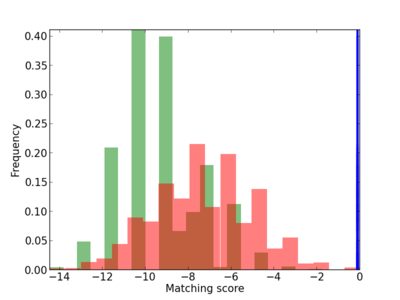 |
7 |
1.0 |
3.30E-04 |
TBX1,
TAB1,
RXRA,
MIB1,
SOX8,
COL13A1,
VEGFA,
TCF25,
CTHRC1,
QKI |
muscle cell differentiation [BP],
cochlea morphogenesis [BP],
heart morphogenesis [BP],
heart development [BP],
coronary artery morphogenesis [BP],
morphogenesis of a branching structure [BP],
in utero embryonic development [BP],
positive regulation of branching involved in ureteric bud morphogenesis [BP],
positive regulation of osteoblast proliferation [BP],
regulation of organ morphogenesis [BP],
lymph vessel development [BP] |
| CG_ffipsc69_promoter_p_RM_12_186_7 |
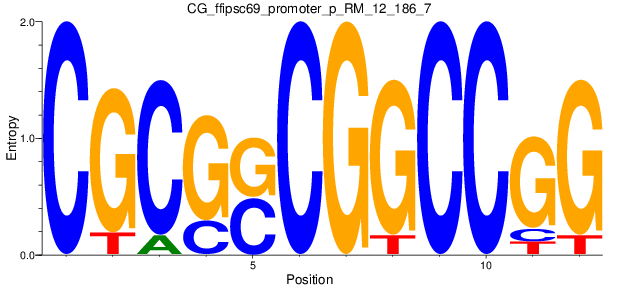 |
 |
2 |
0.4 |
3.26E-04 |
ERI1,
YBX1 |
histone pre-mRNA 3'end processing complex [CC] |
| CG_ffipsc69_promoter_p_RM_12_170_9 |
 |
 |
5 |
0.8 |
5.42E-04 |
GNAS,
HS3ST2 |
[heparan sulfate]-glucosamine 3-sulfotransferase 2 activity [MF],
post-embryonic body morphogenesis [BP] |
| CG_ffipsc69_promoter_p_RM_14_11_3 |
 |
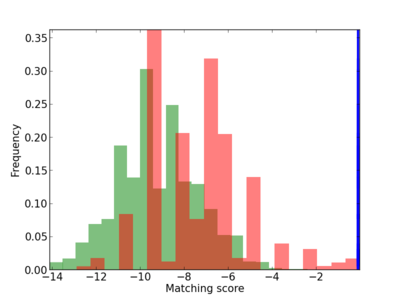 |
2 |
0.4 |
2.64E-04 |
CYR61,
NAGK,
CRTAP,
CCKBR |
peptidyl-proline hydroxylation to 3-hydroxy-L-proline [BP],
gastrin receptor activity [MF],
N-acetylglucosamine kinase activity [MF],
intussusceptive angiogenesis [BP],
type B gastrin/cholecystokinin receptor binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_30_8 |
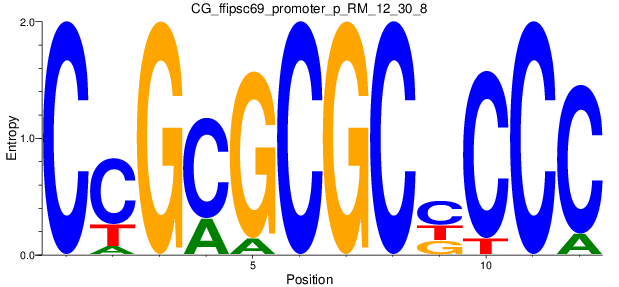 |
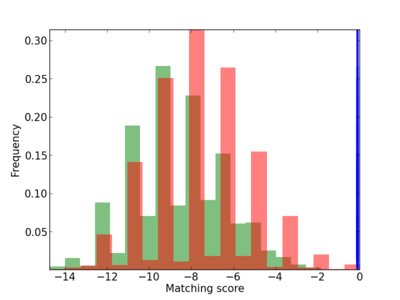 |
5 |
1.0 |
2.70E-04 |
CNTNAP1,
OR1F1,
MAVS,
HSP90AB1,
GABRA5 |
positive regulation of protein import into nucleus, translocation [BP],
signal transduction [BP] |
| CG_ffipsc69_promoter_p_RM_12_269_4 |
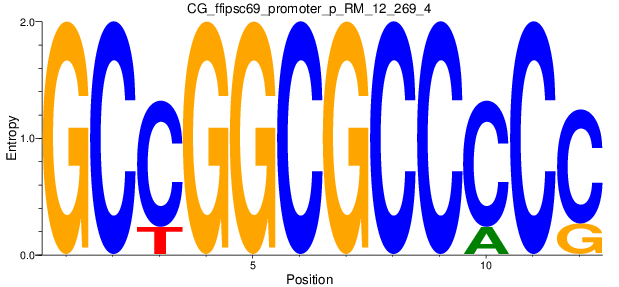 |
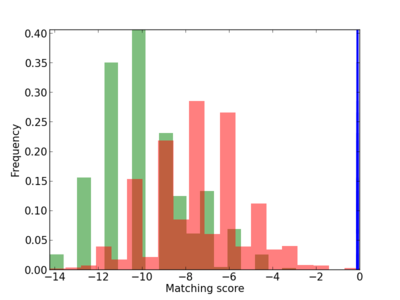 |
3 |
0.4 |
2.12E-04 |
ZRANB1,
NKX2-3 |
protein deubiquitination involved in ubiquitin-dependent protein catabolic process [BP],
post-embryonic digestive tract morphogenesis [BP] |
| CG_ffipsc69_promoter_p_RM_18_1_6 |
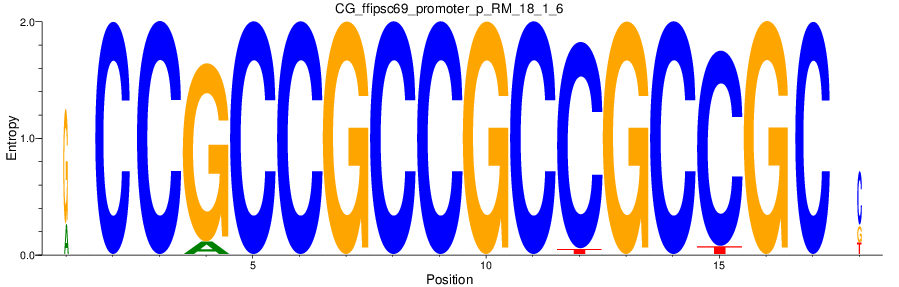 |
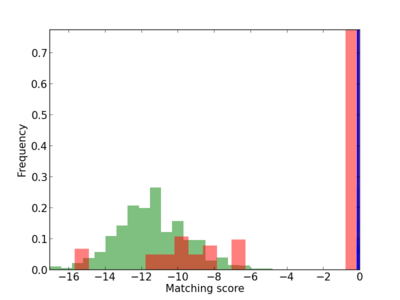 |
4 |
0.4 |
4.92E-04 |
BMPR2,
PKNOX1,
GNAS,
ACVR2A |
positive regulation of osteoblast differentiation [BP],
positive regulation of pathway-restricted SMAD protein phosphorylation [BP],
activin-activated receptor activity [MF],
transforming growth factor beta-activated receptor activity [MF],
transmembrane receptor protein serine/threonine kinase activity [MF],
erythrocyte differentiation [BP] |
| CG_ffipsc69_promoter_p_RM_12_178_6 |
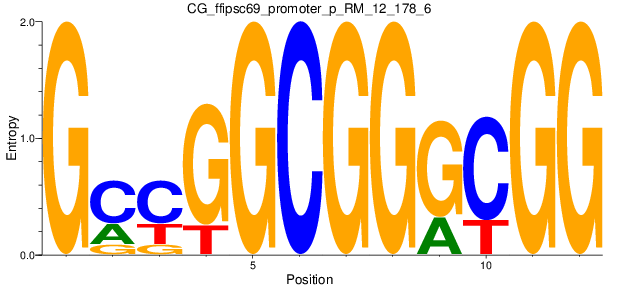 |
 |
5 |
0.8 |
2.69E-04 |
SCIN,
VIL1 |
actin nucleation [BP],
actin filament severing [BP] |
| CG_ffipsc69_promoter_p_RM_12_215_4 |
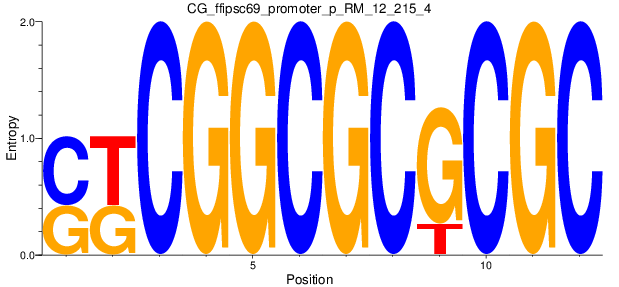 |
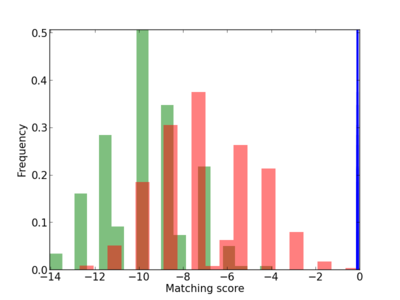 |
5 |
0.8 |
1.96E-04 |
ADCYAP1R1,
TGM4,
EIF4EBP2,
FLVCR1 |
heme export [BP],
cAMP-mediated signaling [BP],
protein polyamination [BP] |
| CG_ffipsc69_promoter_p_RM_12_239_5 |
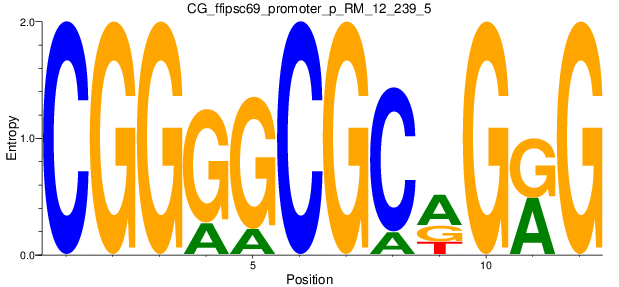 |
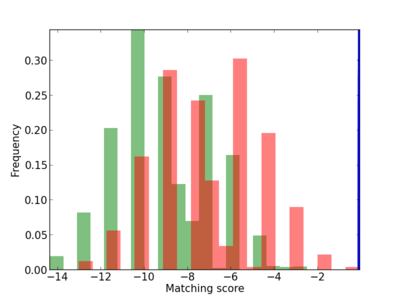 |
3 |
0.6 |
2.68E-04 |
E2F3,
JARID2,
REL,
AGK,
HEY2,
PHF20,
GCDH,
KAT8 |
histone acetyltransferase activity [MF],
glutaryl-CoA dehydrogenase activity [MF],
MLL1 complex [CC],
histone H4-K16 acetylation [BP],
negative regulation of cardiac vascular smooth muscle cell differentiation [BP],
nucleoplasm [CC],
histone acetyltransferase activity (H4-K16 specific) [MF],
histone acetyltransferase activity (H4-K5 specific) [MF],
histone acetyltransferase activity (H4-K8 specific) [MF],
histone H4-K8 acetylation [BP],
histone methyltransferase complex [CC],
acylglycerol kinase activity [MF],
histone H4-K5 acetylation [BP] |
| CG_ffipsc69_promoter_p_RM_12_137_8 |
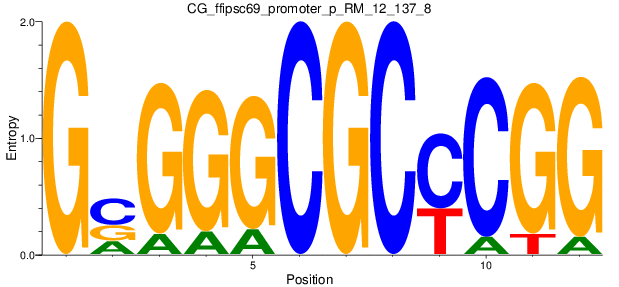 |
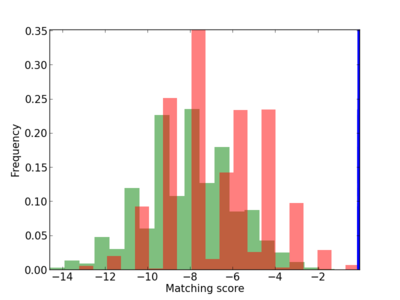 |
5 |
1.0 |
2.23E-04 |
PAX2 |
optic cup morphogenesis involved in camera-type eye development [BP],
positive regulation of optic nerve formation [BP],
optic chiasma development [BP],
vestibulocochlear nerve formation [BP] |
| CG_ffipsc69_promoter_p_RM_12_127_2 |
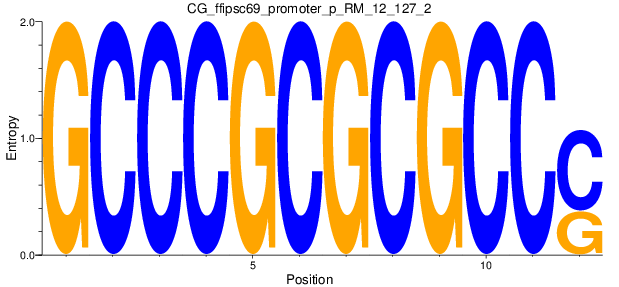 |
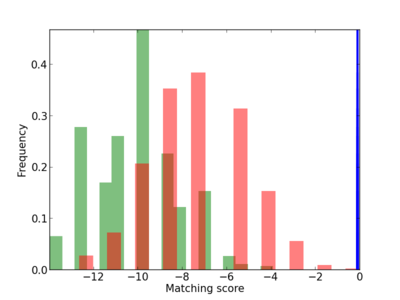 |
4 |
0.8 |
2.51E-04 |
TNFSF11,
TRAF5,
DOPEY2,
SCAMP1,
AMH |
protein transport [BP],
positive regulation of NF-kappaB transcription factor activity [BP] |
| CG_ffipsc69_promoter_p_RM_12_65_5 |
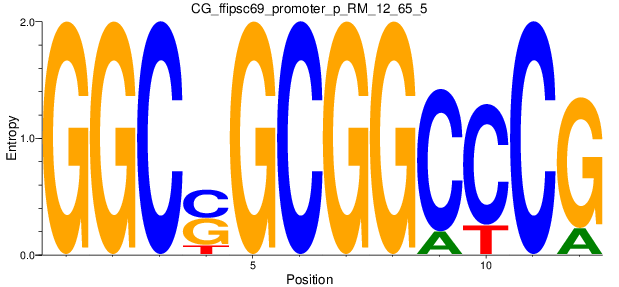 |
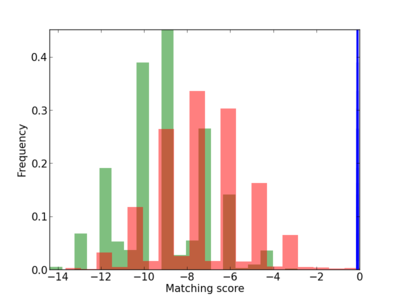 |
6 |
1.0 |
4.04E-04 |
MLL,
PTEN |
negative regulation of cyclin-dependent protein kinase activity involved in G1/S [BP],
postsynaptic density assembly [BP],
positive regulation of cellular response to drug [BP],
negative regulation of ribosome biogenesis [BP],
negative regulation of synaptic vesicle clustering [BP] |
| CG_ffipsc69_promoter_p_RM_14_17_3 |
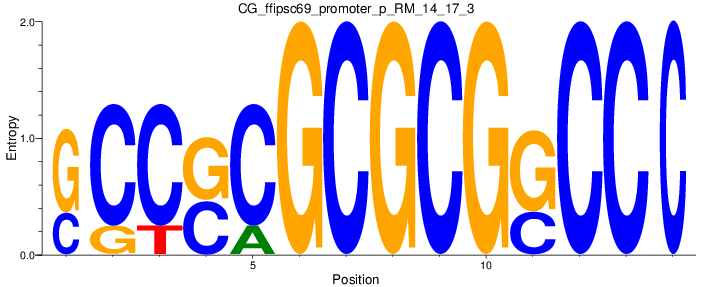 |
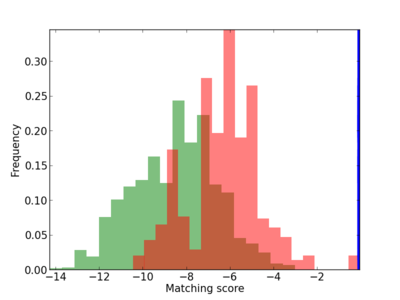 |
2 |
0.4 |
2.36E-04 |
TNFAIP3,
BICD1 |
negative regulation of CD40 signaling pathway [BP],
negative regulation of toll-like receptor 5 signaling pathway [BP],
negative regulation of phospholipase C activity [BP],
host cell viral assembly compartment [CC],
proteinase activated receptor binding [MF],
negative regulation of phospholipase C-activating G-protein coupled receptor signaling pathway [BP],
negative regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway [BP],
regulation of proteinase activated receptor activity [BP],
tolerance induction to lipopolysaccharide [BP],
negative regulation of nucleotide-binding oligomerization domain containing 2 signaling pathway [BP] |
| CG_ffipsc69_promoter_p_RM_12_211_5 |
 |
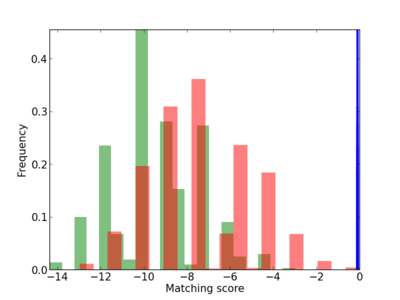 |
7 |
1.0 |
2.30E-04 |
BICD1 |
proteinase activated receptor binding [MF] |
| CG_ffipsc69_promoter_p_RM_14_42_3 |
 |
 |
3 |
0.6 |
2.78E-04 |
SPAG6 |
axoneme [CC] |
| CG_ffipsc69_promoter_p_RM_12_146_8 |
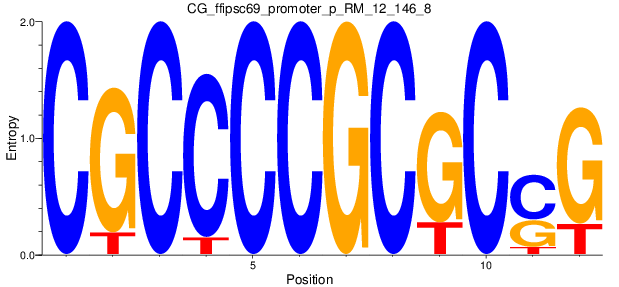 |
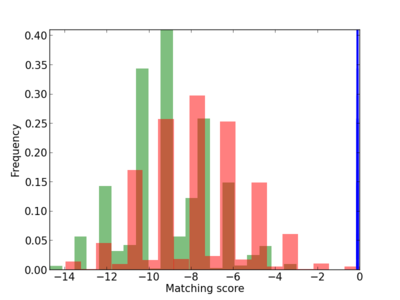 |
3 |
0.6 |
3.83E-04 |
CAD |
dihydroorotase activity [MF],
aspartate binding [MF],
aspartate carbamoyltransferase activity [MF],
carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_258_11 |
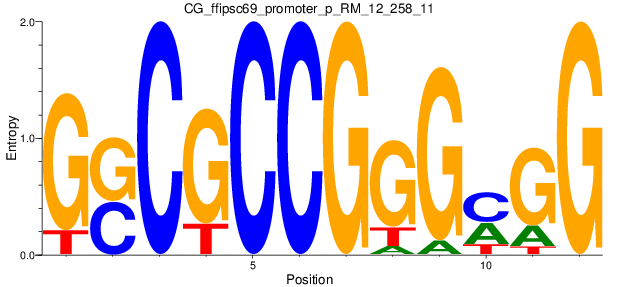 |
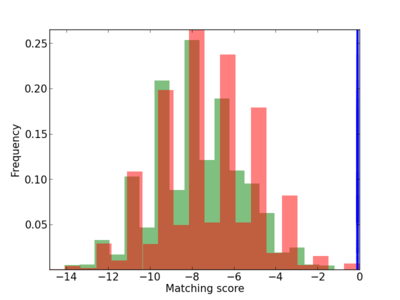 |
5 |
1.0 |
3.83E-04 |
SLC2A4,
PFKFB2,
MGAT2,
PPARD |
linoleic acid binding [MF],
amylopectin biosynthetic process [BP],
alpha-1,6-mannosylglycoprotein 2-beta-N-acetylglucosaminyltransferase activity [MF],
carbohydrate metabolic process [BP] |
| CG_ffipsc69_promoter_p_RM_12_100_4 |
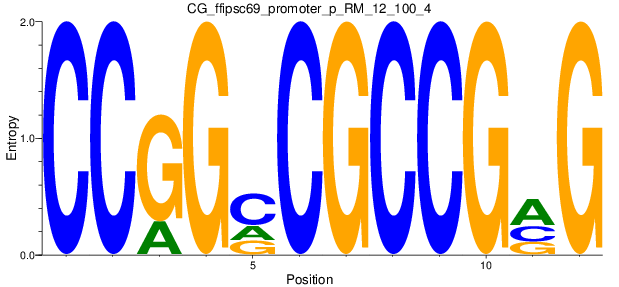 |
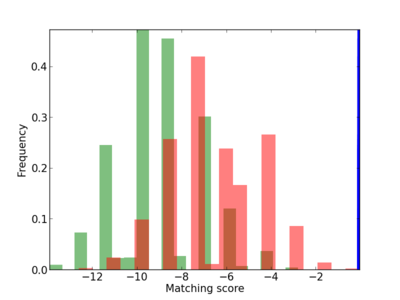 |
1 |
0.2 |
2.32E-04 |
TBC1D25,
B3GALT6 |
galactosylxylosylprotein 3-beta-galactosyltransferase activity [MF],
regulation of autophagic vacuole maturation [BP] |
| CG_ffipsc69_promoter_p_RM_12_36_3 |
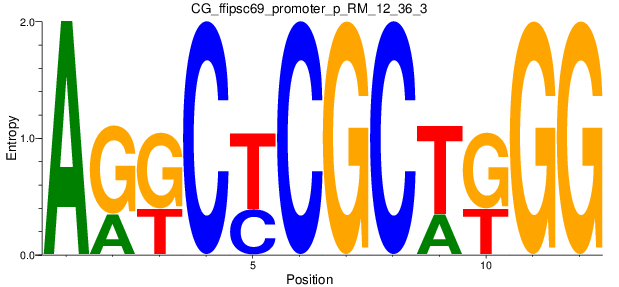 |
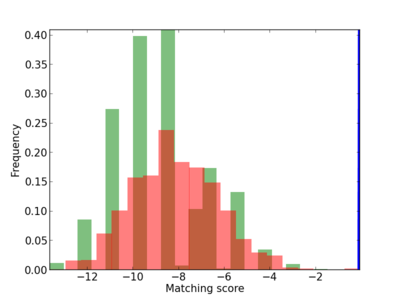 |
1 |
0.2 |
3.30E-04 |
AMDHD2,
PAK2,
GATA4 |
lung lobe formation [BP],
negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis [BP],
N-acetylglucosamine-6-phosphate deacetylase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_67_17 |
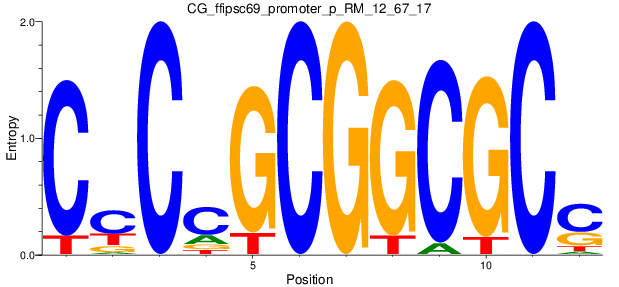 |
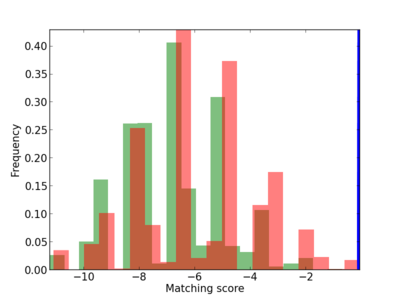 |
5 |
0.8 |
1.98E-04 |
SGOL2,
MTRR,
ACER3 |
phytoceramidase activity [MF],
[methionine synthase] reductase activity [MF],
phytosphingosine biosynthetic process [BP],
meiotic sister chromatid cohesion, centromeric [BP] |
| CG_ffipsc69_promoter_p_RM_12_139_14 |
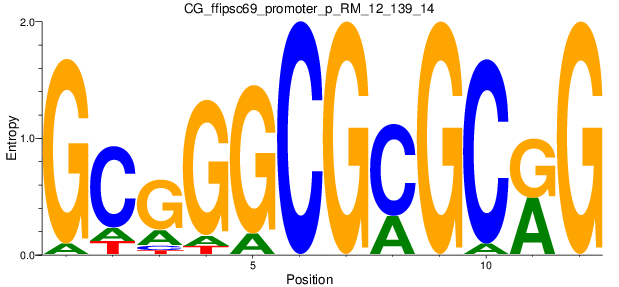 |
 |
5 |
1.0 |
2.39E-04 |
RNF8,
RING1,
L3MBTL1 |
histone binding [MF],
histone H2A ubiquitination [BP],
chromatin modification [BP] |
| CG_ffipsc69_promoter_p_RM_12_174_7 |
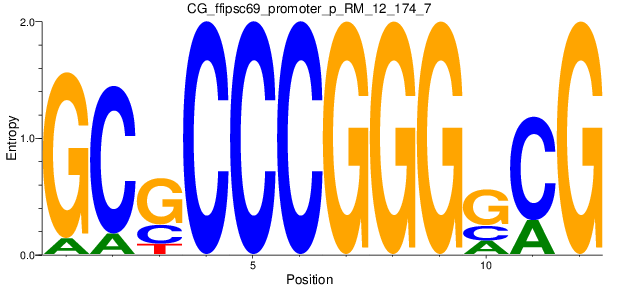 |
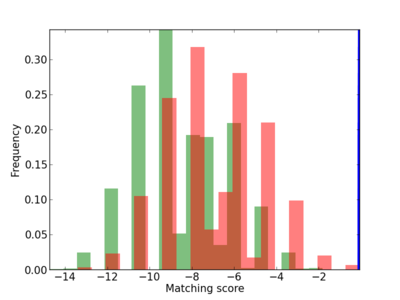 |
3 |
0.6 |
2.75E-04 |
GABARAPL2,
MAP1LC3B |
autophagic vacuole membrane [CC] |
| CG_ffipsc69_promoter_p_RM_12_168_9 |
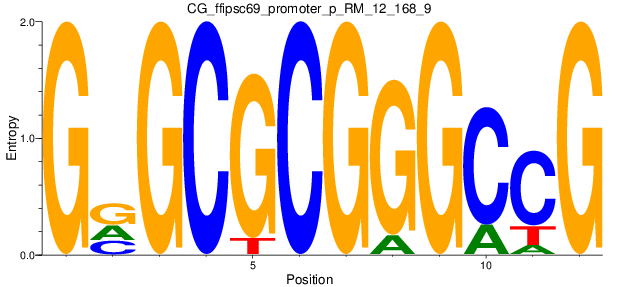 |
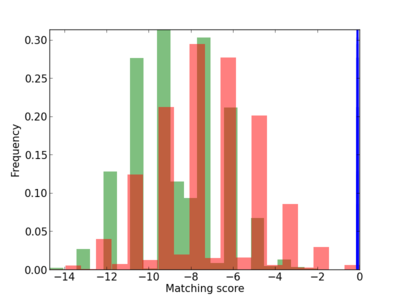 |
5 |
0.8 |
2.78E-04 |
OPRL1 |
nociceptin receptor activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_38_16 |
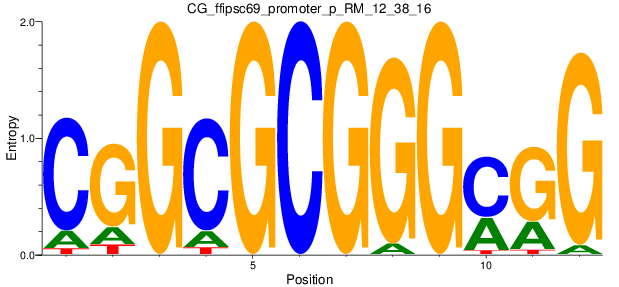 |
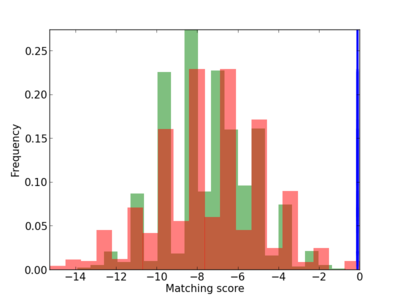 |
5 |
1.0 |
3.40E-04 |
RAC1,
GNAS,
CAMK4,
COL13A1,
SSH3,
MDM2,
GNAI2,
CCNK,
SPATA13 |
regulation of lamellipodium assembly [BP],
extrinsic to plasma membrane [CC],
G-protein beta/gamma-subunit complex binding [MF],
lamellipodium assembly [BP],
endochondral ossification [BP],
positive regulation of protein export from nucleus [BP],
negative regulation of cell cycle arrest [BP] |
| CG_ffipsc69_promoter_p_RM_18_5_1 |
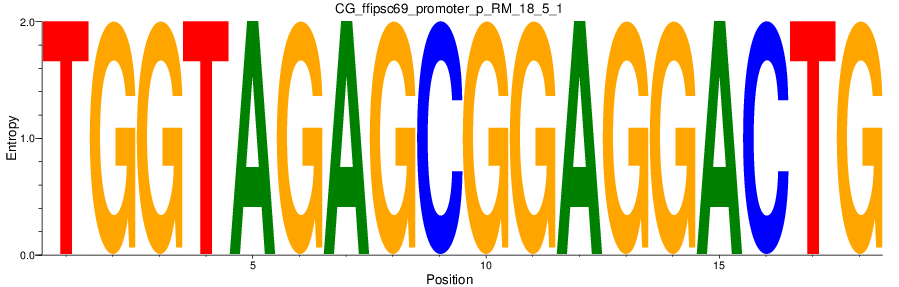 |
 |
1 |
0.2 |
9.75E-04 |
EMILIN1,
ANG |
angiogenin-PRI complex [CC],
extracellular matrix constituent conferring elasticity [MF] |
| CG_ffipsc69_promoter_p_RM_12_125_5 |
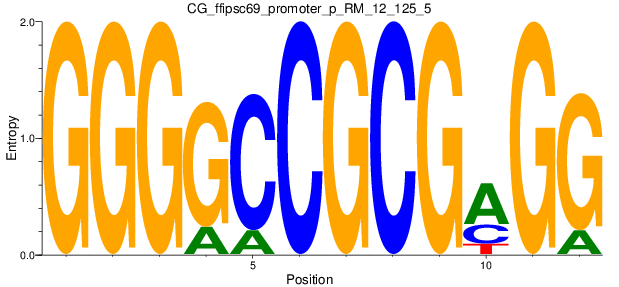 |
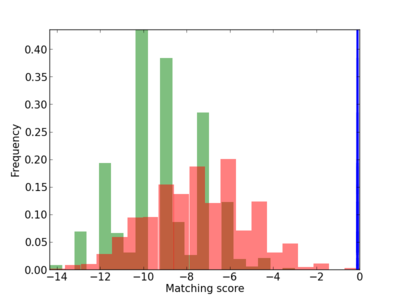 |
5 |
1.0 |
2.87E-04 |
HIC1,
MLLT10,
INSM1,
ZNF276,
PAX2,
GEN1,
ENGASE,
SAP30 |
mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase activity [MF],
positive regulation of optic nerve formation [BP],
optic chiasma development [BP],
DNA binding [MF],
vestibulocochlear nerve formation [BP] |
| CG_ffipsc69_promoter_p_RM_14_12_2 |
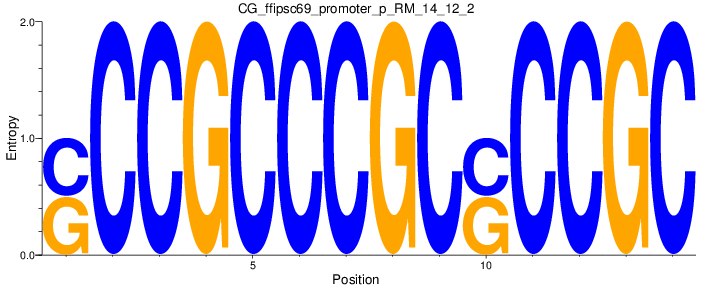 |
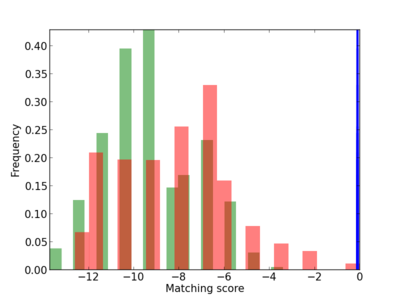 |
3 |
0.4 |
9.83E-05 |
ATP2A2 |
calcium-transporting ATPase activity involved in regulation of cardiac muscle cell membrane potential [MF] |
| CG_ffipsc69_promoter_p_RM_12_217_9 |
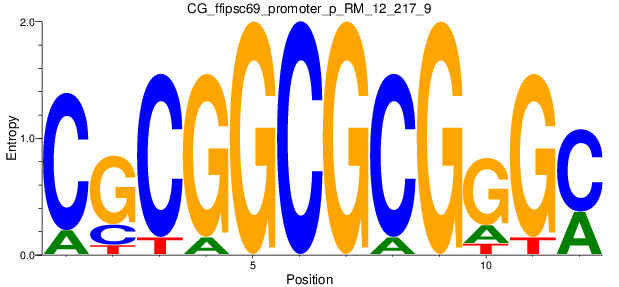 |
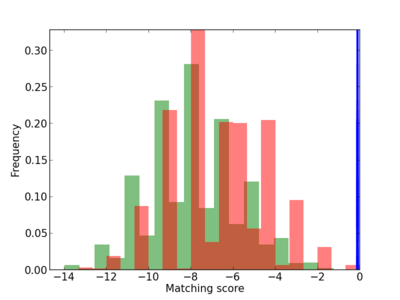 |
4 |
0.8 |
2.49E-04 |
ICT1 |
mitochondrial translational termination [BP],
translation release factor activity, codon nonspecific [MF] |
| CG_ffipsc69_promoter_p_RM_14_23_2 |
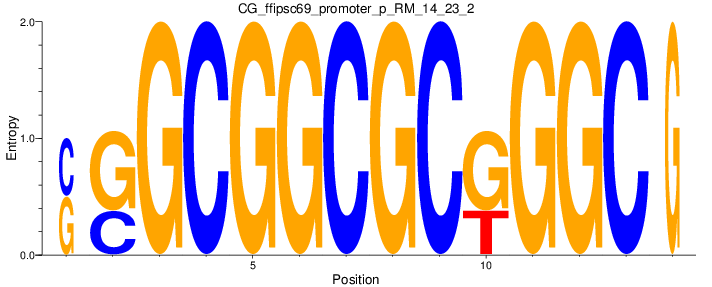 |
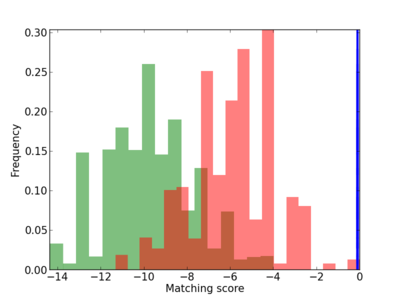 |
2 |
0.4 |
2.13E-04 |
DYNLL2 |
synaptic target recognition [BP] |
| CG_ffipsc69_promoter_p_RM_14_50_3 |
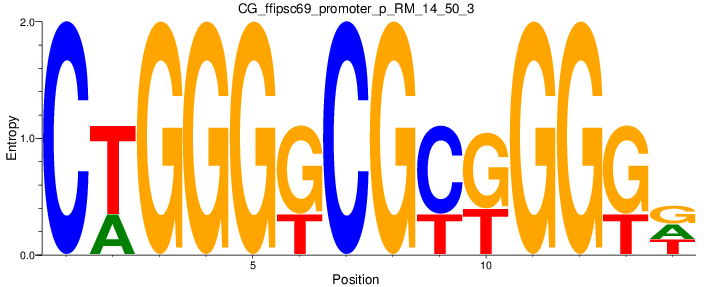 |
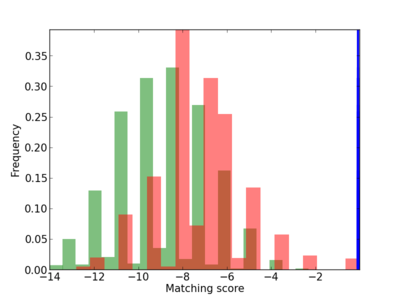 |
5 |
0.8 |
2.90E-04 |
TMX1,
CYP1A2 |
oxidative deethylation [BP],
toxin biosynthetic process [BP],
arsenate reductase (thioredoxin) activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_242_8 |
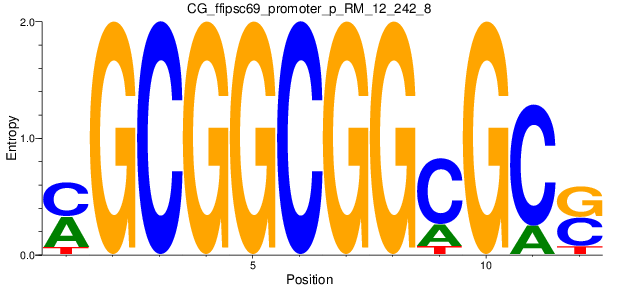 |
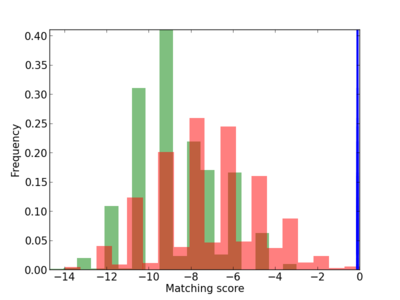 |
4 |
0.8 |
4.34E-04 |
SMARCB1,
CDC73,
ATF7IP,
FOXO3,
MAVS,
YAP1,
BMP6 |
positive regulation of transcription from RNA polymerase II promoter [BP],
positive regulation of transcription, DNA-dependent [BP] |
| CG_ffipsc69_promoter_p_RM_12_94_7 |
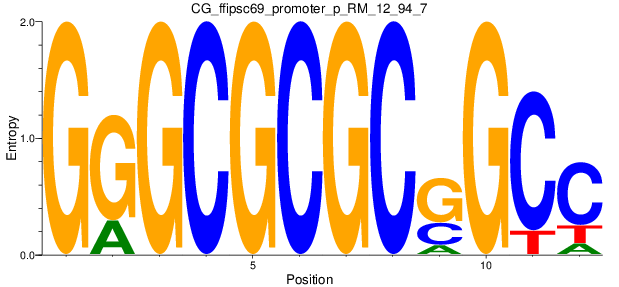 |
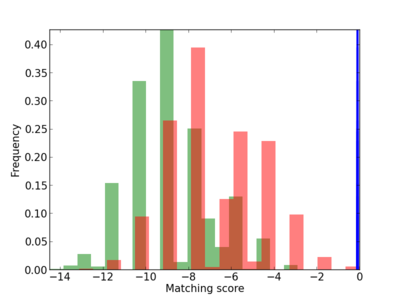 |
8 |
1.0 |
3.94E-04 |
EGR1,
ARID5A,
PRDM8,
NOG,
NTRK1,
AP2A1,
AACS,
KDM3A,
ZNF76,
RBM15B,
CDC42,
CACNA1H |
transcription, DNA-dependent [BP],
axon guidance [BP],
response to electrical stimulus [BP],
response to ethanol [BP],
nervous system development [BP] |
| CG_ffipsc69_promoter_p_RM_12_270_7 |
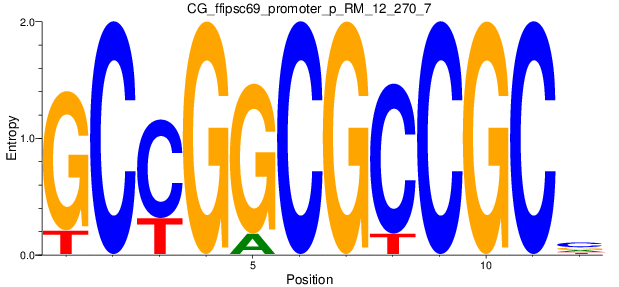 |
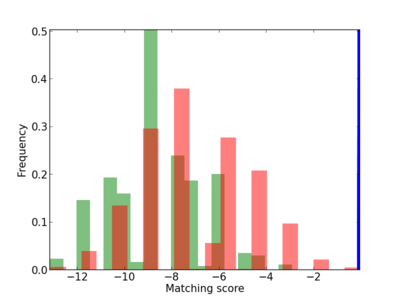 |
3 |
0.6 |
1.79E-04 |
HINFP,
ID2 |
regulation of transcription involved in G1/S phase of mitotic cell cycle [BP],
olfactory bulb development [BP] |
| CG_ffipsc69_promoter_p_RM_12_96_7 |
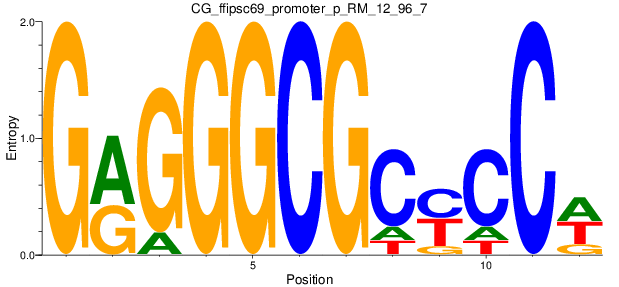 |
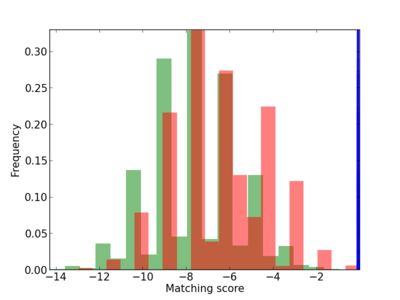 |
2 |
0.4 |
1.96E-04 |
ADRA2C,
MT3 |
activation of protein kinase B activity [BP] |
| CG_ffipsc69_promoter_p_RM_12_202_9 |
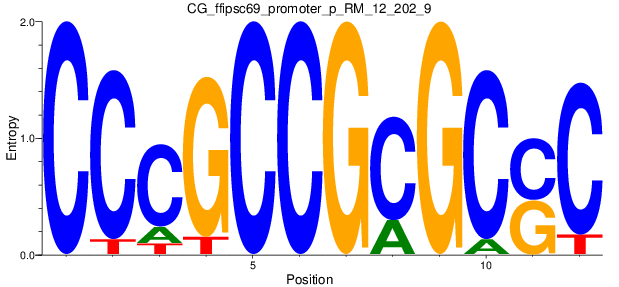 |
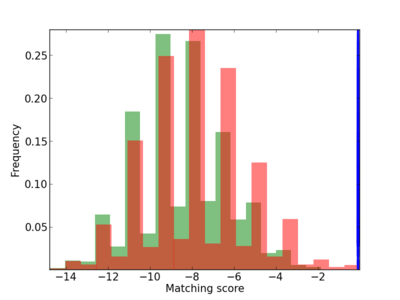 |
7 |
1.0 |
3.11E-04 |
GAA,
BICD1 |
diaphragm contraction [BP],
sucrose metabolic process [BP],
negative regulation of phospholipase C activity [BP],
host cell viral assembly compartment [CC],
maltose metabolic process [BP],
regulation of proteinase activated receptor activity [BP],
proteinase activated receptor binding [MF],
negative regulation of phospholipase C-activating G-protein coupled receptor signaling pathway [BP],
vacuolar sequestering [BP] |
| CG_ffipsc69_promoter_p_RM_12_284_8 |
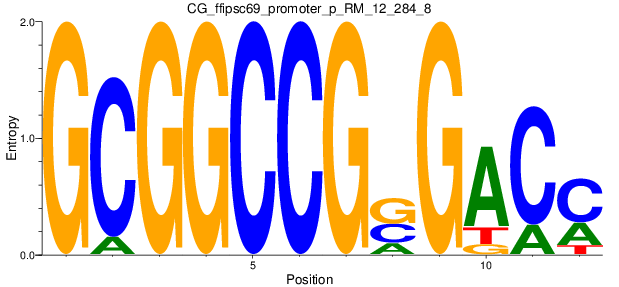 |
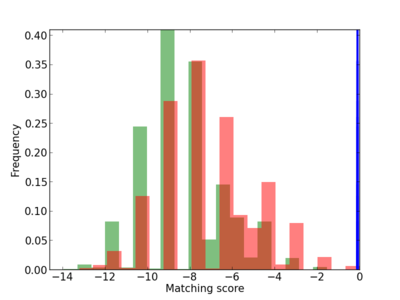 |
5 |
0.8 |
2.90E-04 |
PIN1 |
regulation of cytokinesis [BP] |
| CG_ffipsc69_promoter_p_RM_14_21_3 |
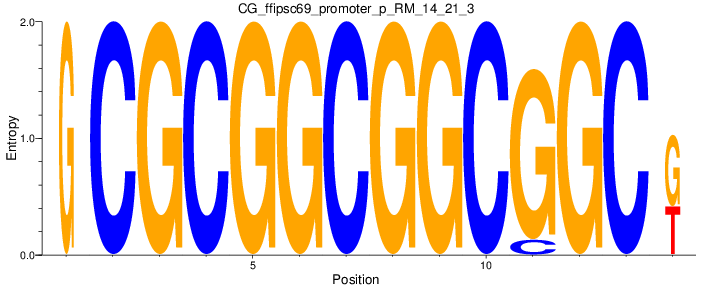 |
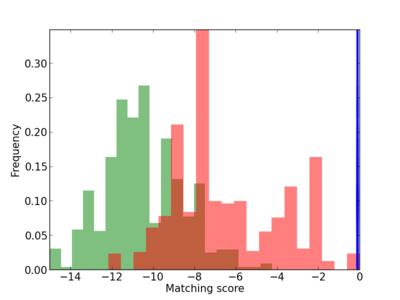 |
13 |
1.0 |
3.01E-04 |
PDE1B,
PGS1,
NPL |
CDP-diacylglycerol-glycerol-3-phosphate 3-phosphatidyltransferase activity [MF],
N-acetylneuraminate lyase activity [MF],
calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_70_6 |
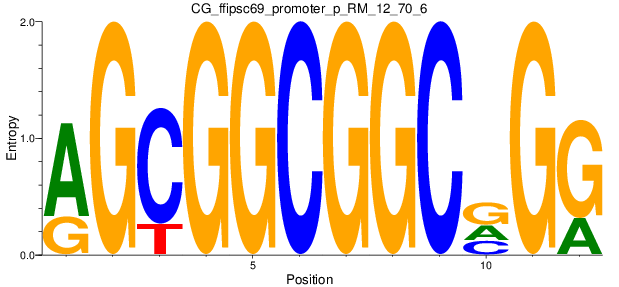 |
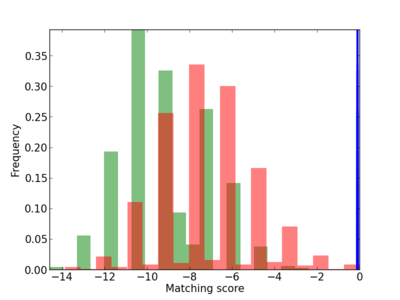 |
4 |
0.8 |
4.29E-04 |
NCMAP,
CNTNAP1 |
paranode region of axon [CC] |
| CG_ffipsc69_promoter_p_RM_14_46_2 |
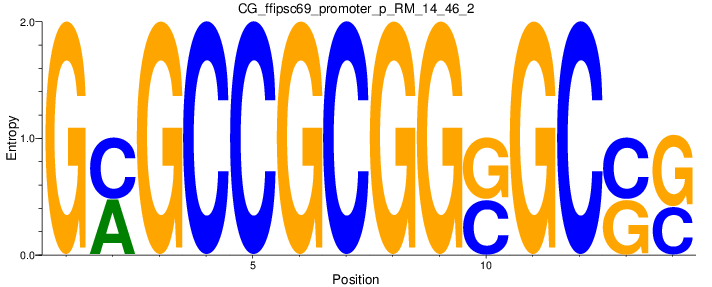 |
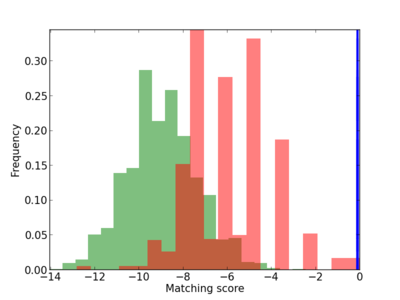 |
5 |
0.8 |
2.43E-04 |
ARPC2 |
AP-1 adaptor complex binding [MF],
Arp2/3 complex binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_85_17 |
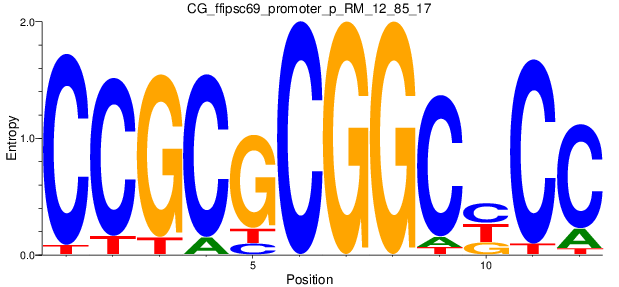 |
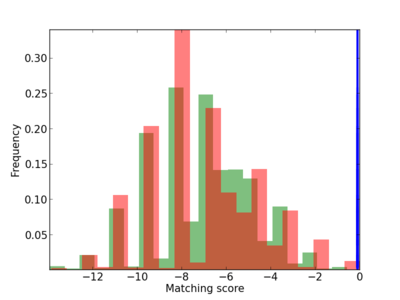 |
4 |
0.8 |
2.34E-04 |
UNC45B,
STUB1 |
Hsp90 protein binding [MF],
ubiquitin conjugating enzyme complex [CC] |
| CG_ffipsc69_promoter_p_RM_12_244_2 |
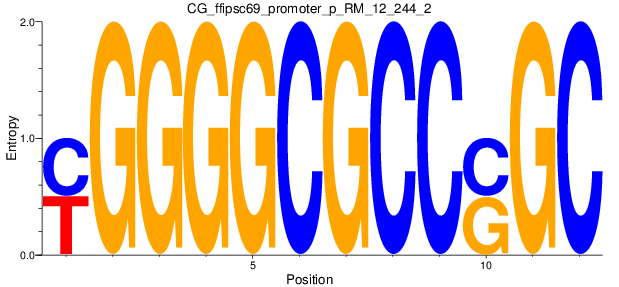 |
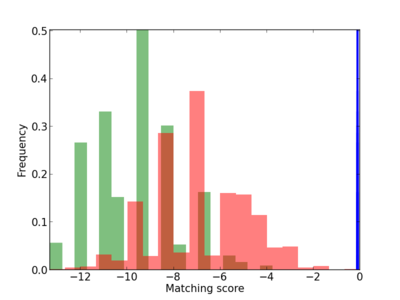 |
4 |
0.8 |
8.03E-05 |
CACNA1C,
CARM1,
NAGS,
EGLN2 |
intracellular estrogen receptor signaling pathway [BP],
histone H3-R17 methylation [BP],
histone methyltransferase activity (H3-R17 specific) [MF],
caveolar macromolecular signaling complex [CC],
acetyl-CoA:L-glutamate N-acetyltransferase activity [MF],
calcium-mediated signaling using extracellular calcium source [BP] |
| CG_ffipsc69_promoter_p_RM_12_209_12 |
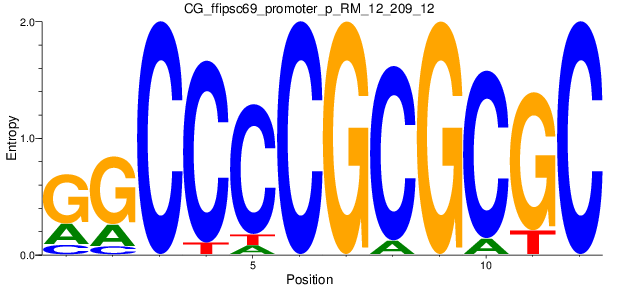 |
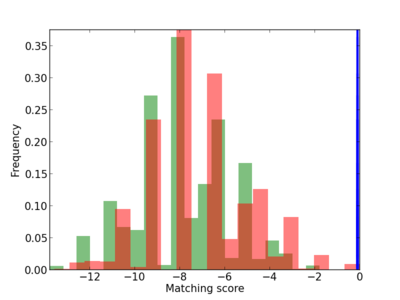 |
6 |
0.6 |
2.13E-04 |
TNFRSF14,
TNFRSF11A,
ADORA2A |
tumor necrosis factor-mediated signaling pathway [BP],
tumor necrosis factor-activated receptor activity [MF],
positive regulation of peptidyl-tyrosine phosphorylation [BP],
negative regulation of alpha-beta T cell activation [BP] |
| CG_ffipsc69_promoter_p_RM_12_118_10 |
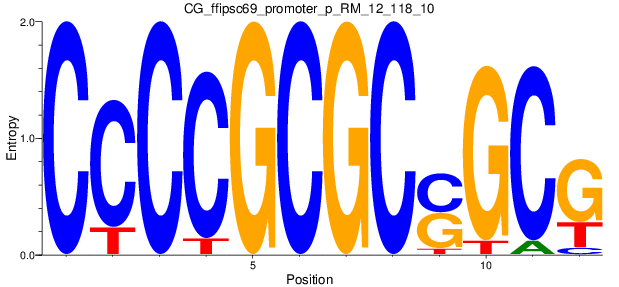 |
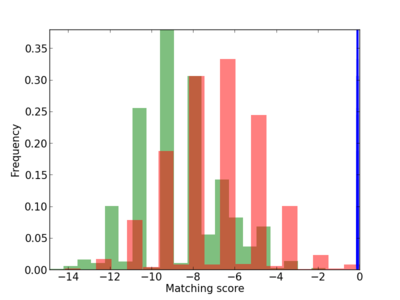 |
2 |
0.4 |
2.41E-04 |
PKN2,
CDC25B |
positive regulation of cell cycle cytokinesis [BP] |
| CG_ffipsc69_promoter_p_RM_12_221_4 |
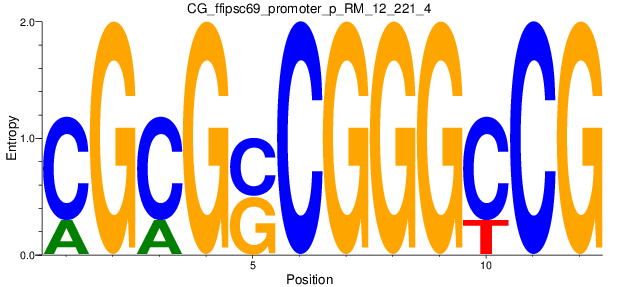 |
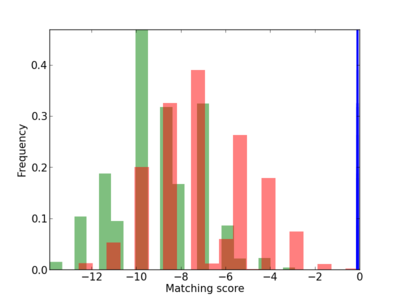 |
3 |
0.6 |
2.57E-04 |
CHAC1,
ATP9B,
L3MBTL1 |
condensed chromosome [CC],
nucleosome binding [MF],
trans-Golgi network [CC] |
| CG_ffipsc69_promoter_p_RM_14_52_3 |
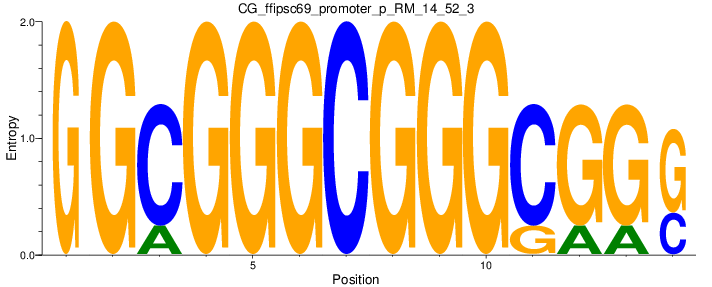 |
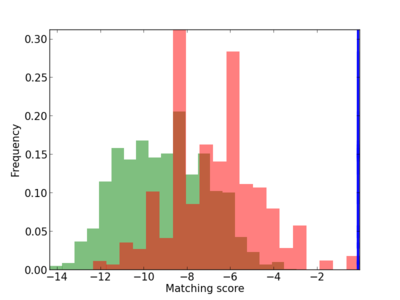 |
5 |
0.8 |
4.66E-04 |
PTPN12,
ABI2 |
SH3 domain binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_78_7 |
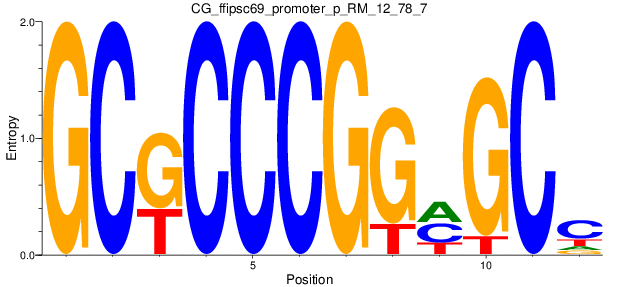 |
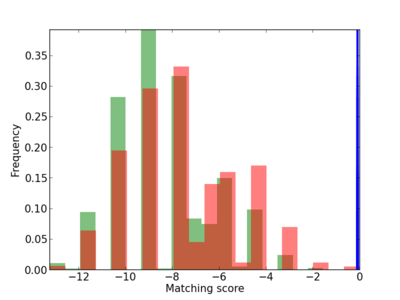 |
4 |
0.8 |
9.55E-05 |
PRKCG |
innervation [BP] |
| CG_ffipsc69_promoter_p_RM_12_254_6 |
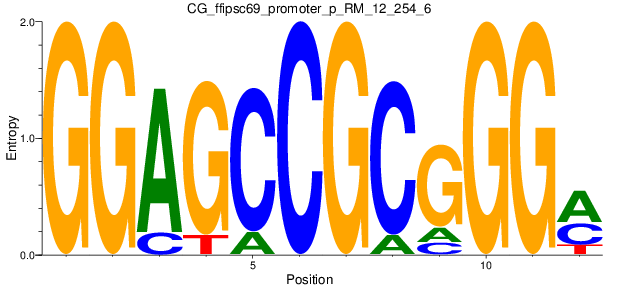 |
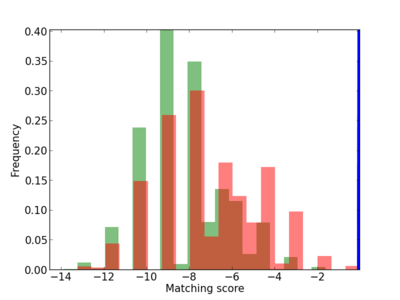 |
4 |
0.8 |
2.65E-04 |
SPHK1 |
sphingoid catabolic process [BP] |
| CG_ffipsc69_promoter_p_RM_12_15_9 |
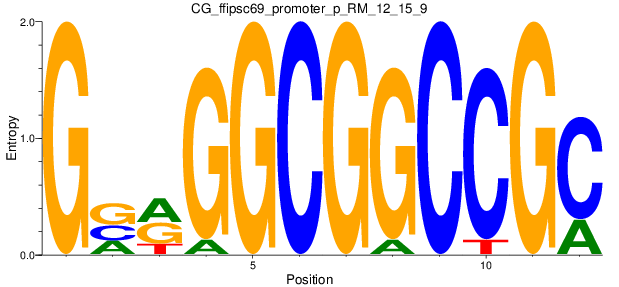 |
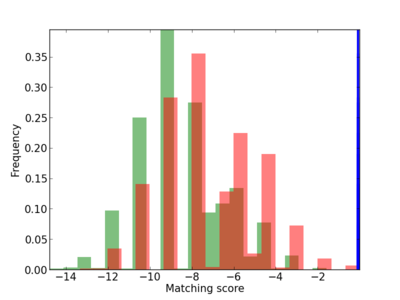 |
2 |
0.4 |
3.72E-04 |
RDH10,
SUFU,
RYR3 |
nose development [BP],
calcium-release channel activity [MF],
negative regulation of transcription factor import into nucleus [BP] |
| CG_ffipsc69_promoter_p_RM_12_114_12 |
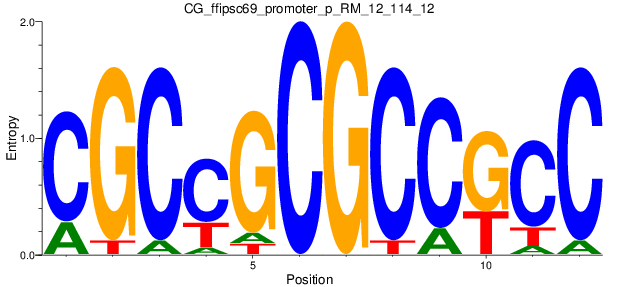 |
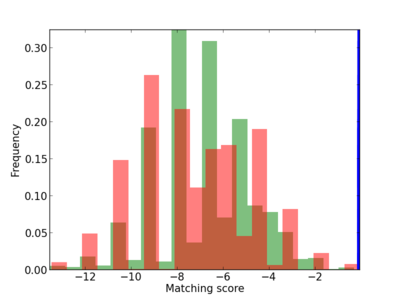 |
6 |
1.0 |
2.17E-04 |
TNFAIP3,
B3GNT5 |
negative regulation of CD40 signaling pathway [BP],
beta-galactosyl-N-acetylglucosaminylgalactosylglucosyl-ceramide beta-1,3-acetylglucosaminyltransferase activity [MF],
lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase activity [MF],
negative regulation of toll-like receptor 5 signaling pathway [BP],
lipopolysaccharide N-acetylglucosaminyltransferase activity [MF],
negative regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway [BP],
tolerance induction to lipopolysaccharide [BP],
negative regulation of nucleotide-binding oligomerization domain containing 2 signaling pathway [BP] |
| CG_ffipsc69_promoter_p_RM_12_189_3 |
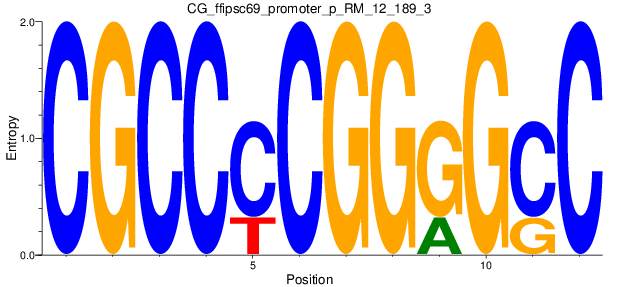 |
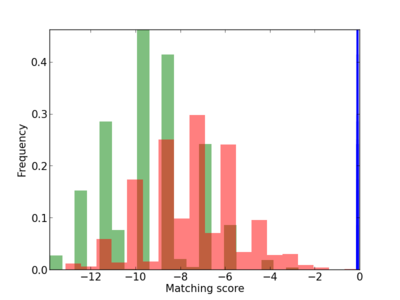 |
5 |
1.0 |
1.71E-04 |
SAT1 |
spermidine binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_153_8 |
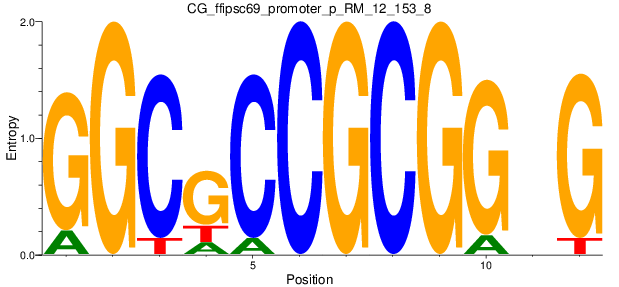 |
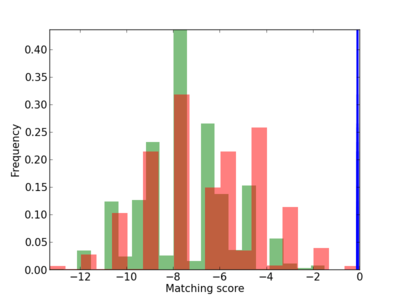 |
6 |
1.0 |
3.73E-04 |
OGDH |
oxoglutarate dehydrogenase (NAD+) activity [MF] |
| CG_ffipsc69_promoter_p_RM_12_66_11 |
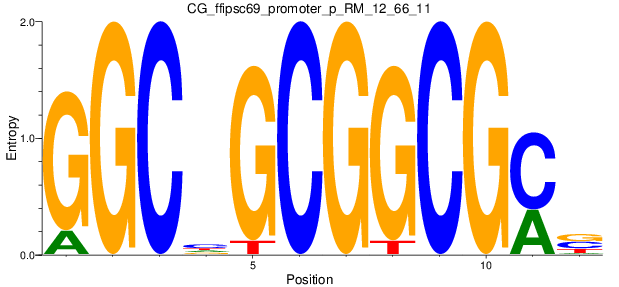 |
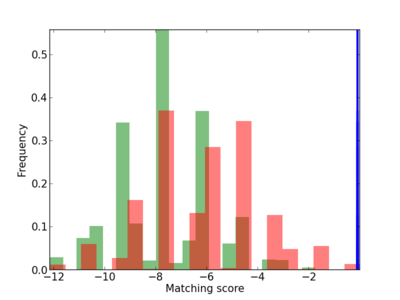 |
6 |
1.0 |
4.91E-04 |
FBXO45,
EN2,
FOS,
PRDM8,
ARID2,
ATF4,
ZNF583,
NPAS1,
INA,
SOX8,
MAFK,
ZNF584,
FOSL2,
SOX11,
IKZF1,
CNOT6,
PKNOX2,
MLF1 |
central nervous system development [BP],
oligodendrocyte differentiation [BP],
sequence-specific DNA binding transcription factor activity [MF],
transcription, DNA-dependent [BP],
sequence-specific DNA binding [MF],
nucleic acid binding transcription factor activity [MF],
nervous system development [BP] |
| CG_ffipsc69_promoter_p_RM_12_17_8 |
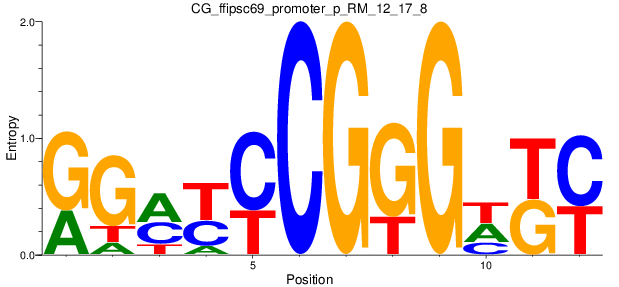 |
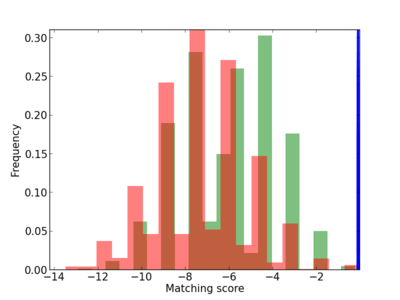 |
4 |
0.6 |
2.52E-04 |
PAX1,
SLC35A1,
LRP3 |
CD4-positive, alpha-beta T cell differentiation [BP],
CMP-N-acetylneuraminate transmembrane transporter activity [MF],
receptor-mediated endocytosis [BP] |
| CG_ffipsc69_promoter_p_RM_12_69_6 |
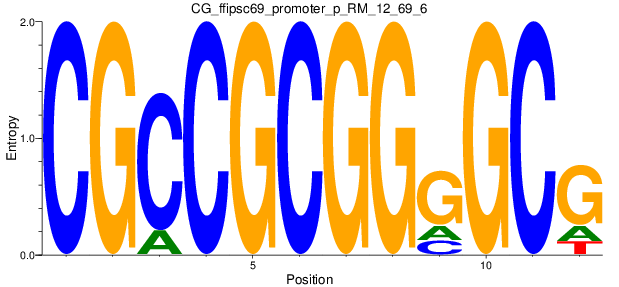 |
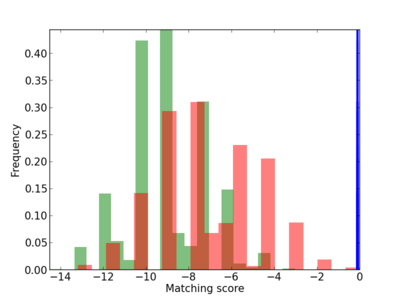 |
5 |
1.0 |
2.05E-04 |
APOC1,
CTR9 |
negative regulation of phosphatidylcholine catabolic process [BP],
positive regulation of histone H3-K79 methylation [BP] |
| CG_ffipsc69_promoter_p_RM_12_219_9 |
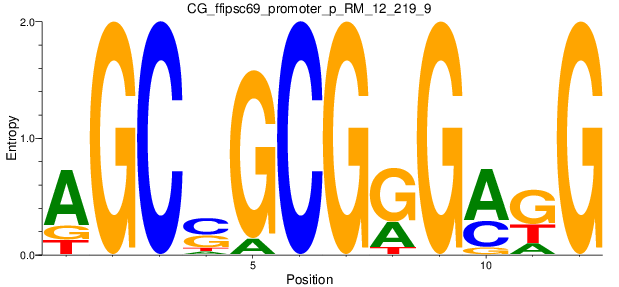 |
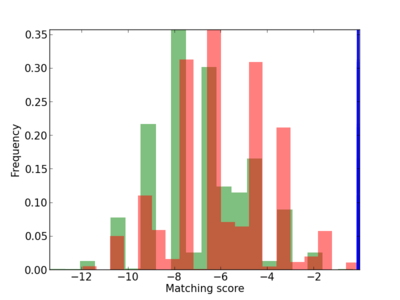 |
4 |
0.8 |
2.60E-04 |
NR2E1,
BAG4,
HELT |
negative regulation of phosphatidylinositol-3,4,5-trisphosphate 5-phosphatase activity [BP],
amygdala development [BP],
GABAergic neuron differentiation in basal ganglia [BP] |
| CG_ffipsc69_promoter_p_RM_12_77_8 |
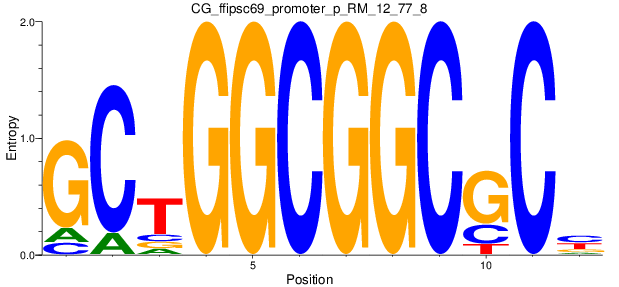 |
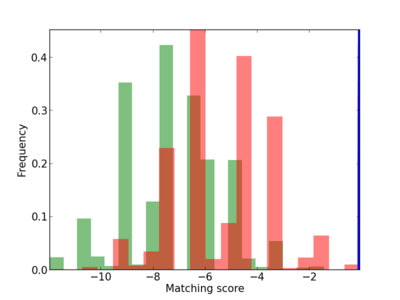 |
5 |
1.0 |
5.16E-04 |
PRRX2,
PINK1 |
embryonic cranial skeleton morphogenesis [BP],
C3HC4-type RING finger domain binding [MF] |
| CG_ffipsc69_promoter_p_RM_12_190_6 |
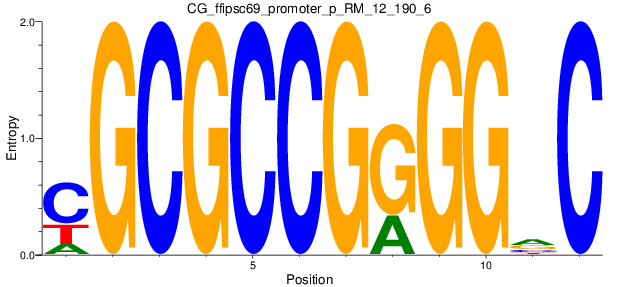 |
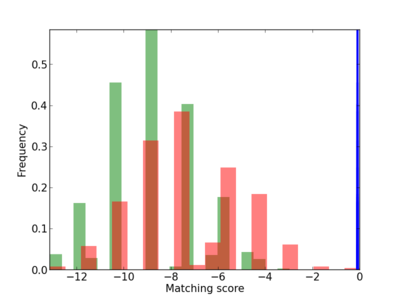 |
5 |
0.8 |
3.62E-04 |
BMPR1A,
ADORA1,
FGF2,
P2RX4,
GABRA5 |
stem cell development [BP],
neuronal cell body [CC],
dendrite [CC],
terminal bouton [CC] |
| CG_ffipsc69_promoter_p_RM_12_227_9 |
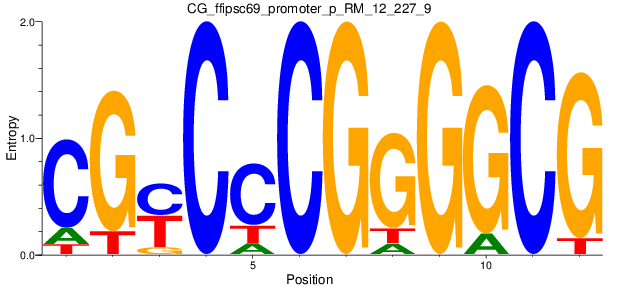 |
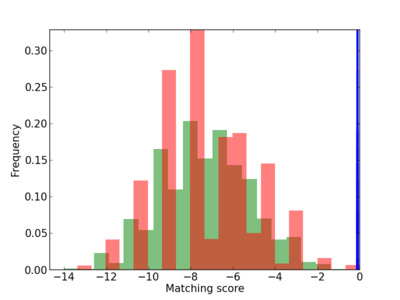 |
4 |
0.6 |
2.73E-04 |
NAGS,
MOV10 |
acetyl-CoA:L-glutamate N-acetyltransferase activity [MF],
mRNA cleavage involved in gene silencing by miRNA [BP] |
| CG_ffipsc69_promoter_p_RM_12_93_10 |
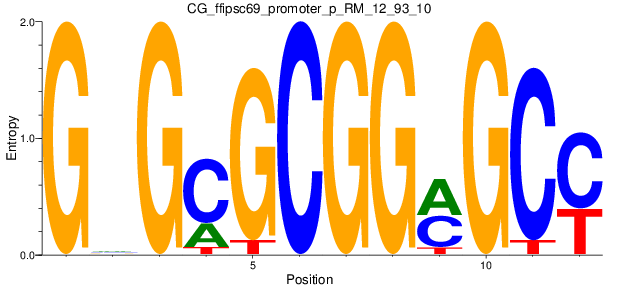 |
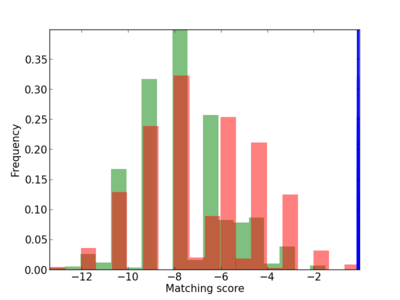 |
5 |
0.8 |
3.08E-04 |
SAC3D1,
PRKCE,
TRIM71,
YBEY,
PKD2,
PLK5,
NR2F2,
PTEN,
CLYBL,
ALKBH5,
FSD1 |
mitosis [BP],
negative regulation of cyclin-dependent protein kinase activity [BP],
metal ion binding [MF],
G1/S transition of mitotic cell cycle [BP],
regulation of cyclin-dependent protein kinase activity [BP],
negative regulation of G1/S transition of mitotic cell cycle [BP] |
| CG_ffipsc69_promoter_p_RM_12_44_5 |
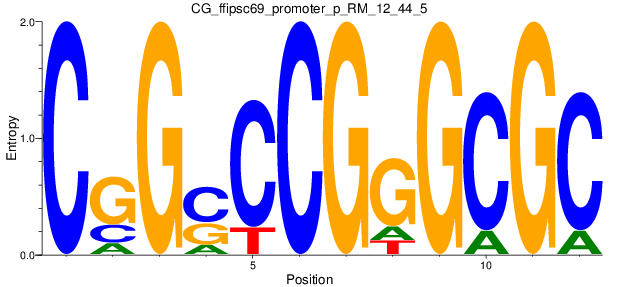 |
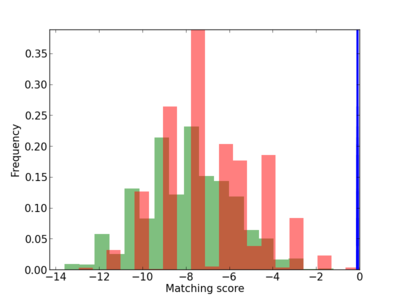 |
2 |
0.4 |
2.42E-04 |
PRKAR2B,
AATF,
ACAT1 |
negative regulation of superoxide anion generation [BP],
response to antipsychotic drug [BP],
metanephric proximal convoluted tubule development [BP] |
| CG_ffipsc69_promoter_p_RM_12_92_7 |
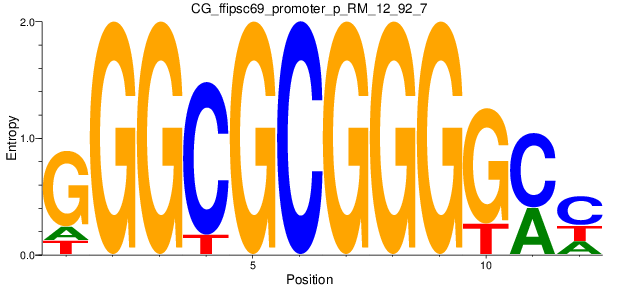 |
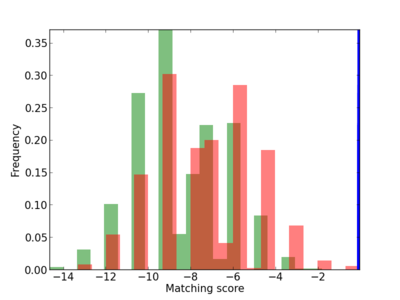 |
3 |
0.6 |
1.88E-04 |
MAP2K1,
B4GALNT2,
TGFBR2 |
negative regulation of cell-cell adhesion [BP],
mitogen-activated protein kinase kinase kinase binding [MF] |
| CG_ffipsc69_promoter_p_RM_14_37_9 |
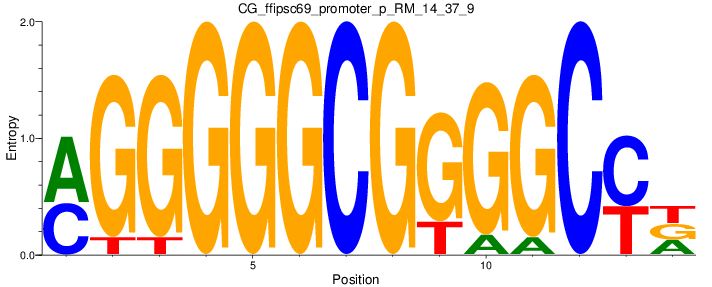 |
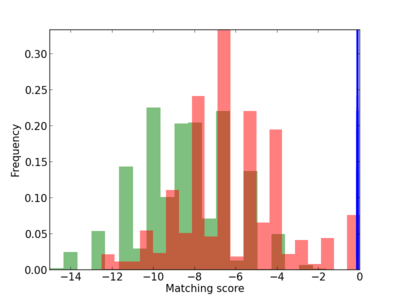 |
9 |
1.0 |
2.90E-04 |
AAMDC,
CREB1,
PIAS3,
PPARD,
TRPM4,
ITPR3,
RYR1 |
positive regulation of fat cell differentiation [BP],
intracellular ligand-gated calcium channel activity [MF],
calcium channel activity [MF],
calcium-release channel activity [MF],
nucleic acid binding [MF] |























































































































































































































































































































































































































































































































































